Show the package imports
import random
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
from keras.models import Sequential
from keras.layers import Dense, Input
from keras.callbacks import EarlyStoppingACTL3143 & ACTL5111 Deep Learning for Actuaries
import random
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
from keras.models import Sequential
from keras.layers import Dense, Input
from keras.callbacks import EarlyStopping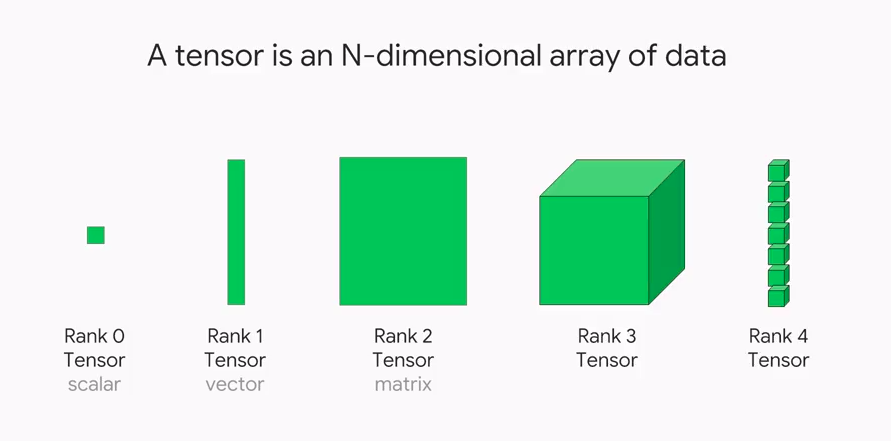
axis argument in numpyStarting with a (3, 4)-shaped matrix:
X = np.arange(12).reshape(3,4)
Xarray([[ 0, 1, 2, 3],
[ 4, 5, 6, 7],
[ 8, 9, 10, 11]])The above code creates an array with values from 0 to 11 and converts that array into a matrix with 3 rows and 4 columns.
axis=0: (3, 4) \leadsto (4,).
X.sum(axis=0)array([12, 15, 18, 21])The above code returns the column sum. This changes the shape of the matrix from (3,4) to (4,). Similarly, X.sum(axis=1) returns row sums and will change the shape of the matrix from (3,4) to (3,).
axis=1: (3, 4) \leadsto (3,).
X.prod(axis=1)array([ 0, 840, 7920])The return value’s rank is one less than the input’s rank.
The axis parameter tells us which dimension is removed.
axis & keepdimsWith keepdims=True, the rank doesn’t change.
X = np.arange(12).reshape(3,4)
Xarray([[ 0, 1, 2, 3],
[ 4, 5, 6, 7],
[ 8, 9, 10, 11]])axis=0: (3, 4) \leadsto (1, 4).
X.sum(axis=0, keepdims=True)array([[12, 15, 18, 21]])axis=1: (3, 4) \leadsto (3, 1).
X.prod(axis=1, keepdims=True)array([[ 0],
[ 840],
[7920]])X / X.sum(axis=1)ValueError: operands could not be broadcast together with shapes (3,4) (3,) X / X.sum(axis=1, keepdims=True)array([[0. , 0.17, 0.33, 0.5 ],
[0.18, 0.23, 0.27, 0.32],
[0.21, 0.24, 0.26, 0.29]])Say we had n observations of a time series x_1, x_2, \dots, x_n.
This \mathbf{x} = (x_1, \dots, x_n) would have shape (n,) & rank 1.
If instead we had a batch of b time series’
\mathbf{X} = \begin{pmatrix} x_7 & x_8 & \dots & x_{7+n-1} \\ x_2 & x_3 & \dots & x_{2+n-1} \\ \vdots & \vdots & \ddots & \vdots \\ x_3 & x_4 & \dots & x_{3+n-1} \\ \end{pmatrix} \,,
the batch \mathbf{X} would have shape (b, n) & rank 2.
Multivariate time series consists of more than 1 variable observation at a given time point. Following example has two variables x and y.
| t | x | y |
|---|---|---|
| 0 | x_0 | y_0 |
| 1 | x_1 | y_1 |
| 2 | x_2 | y_2 |
| 3 | x_3 | y_3 |
Say n observations of the m time series, would be a shape (n, m) matrix of rank 2.
In Keras, a batch of b of these time series has shape (b, n, m) and has rank 3.
Use \mathbf{x}_t \in \mathbb{R}^{1 \times m} to denote the vector of all time series at time t. Here, \mathbf{x}_t = (x_t, y_t).
A recurrence relation is an equation that expresses each element of a sequence as a function of the preceding ones. More precisely, in the case where only the immediately preceding element is involved, a recurrence relation has the form
u_n = \psi(n, u_{n-1}) \quad \text{ for } \quad n > 0.
Example: Factorial n! = n (n-1)! for n > 0 given 0! = 1.
The RNN processes each data in the sequence one by one, while keeping memory of what came before.
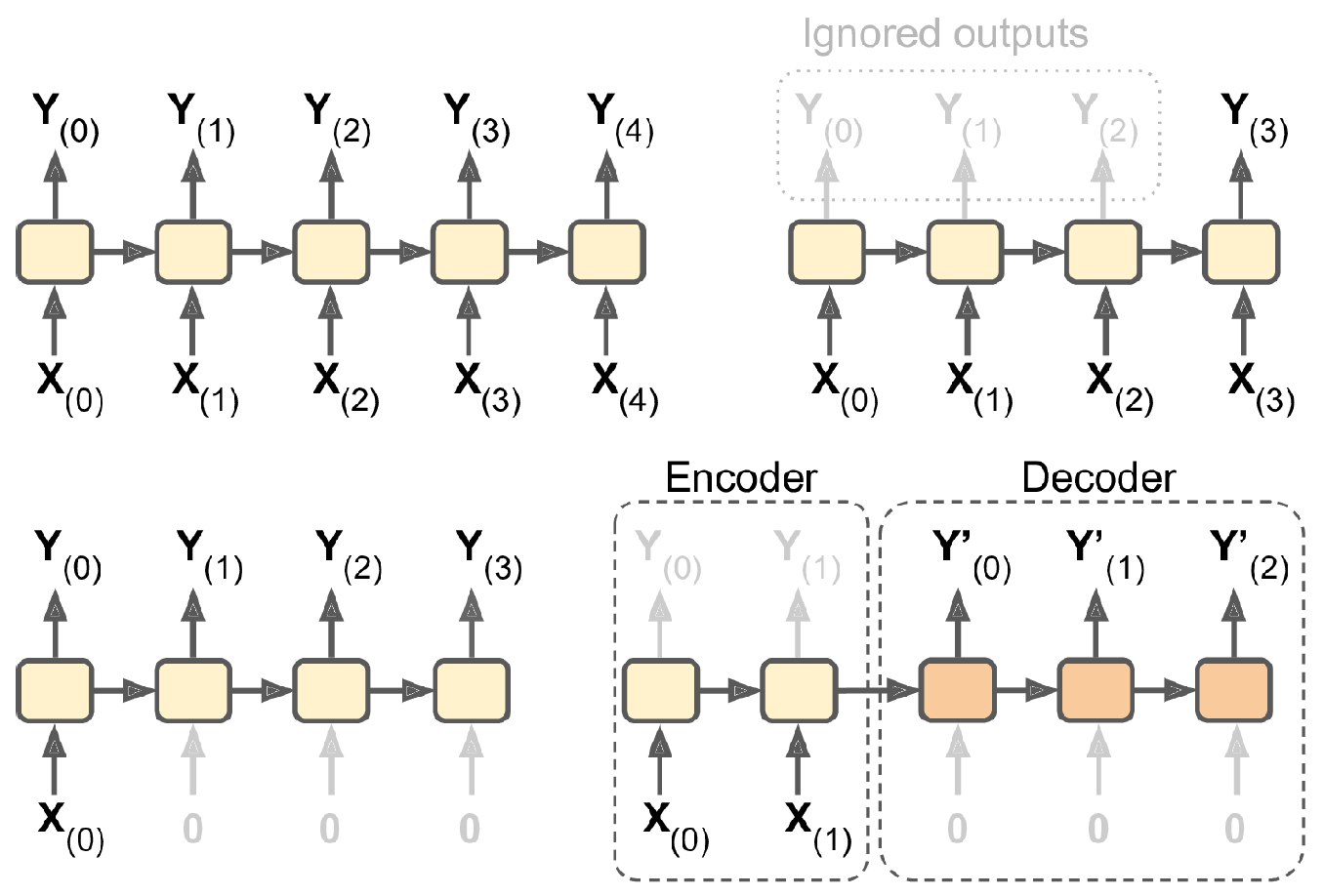
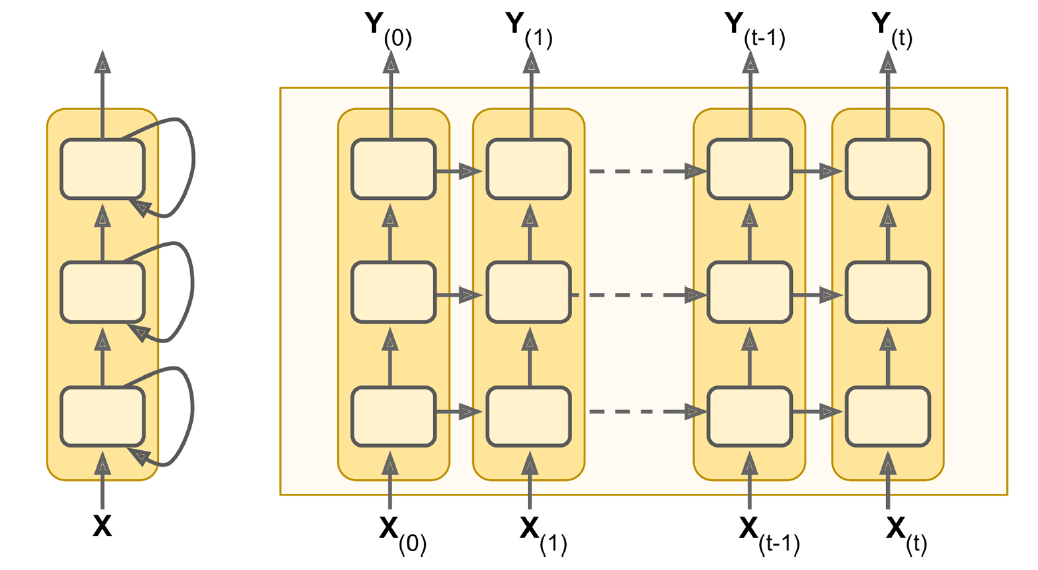
The following figure shows how the recurrent neural network combines an input X_l with a preprocessed state of the process A_l to produce the output O_l. RNNs have a cyclic information processing structure that enables them to pass information sequentially from previous inputs. RNNs can capture dependencies and patterns in sequential data, making them useful for analysing time series data.
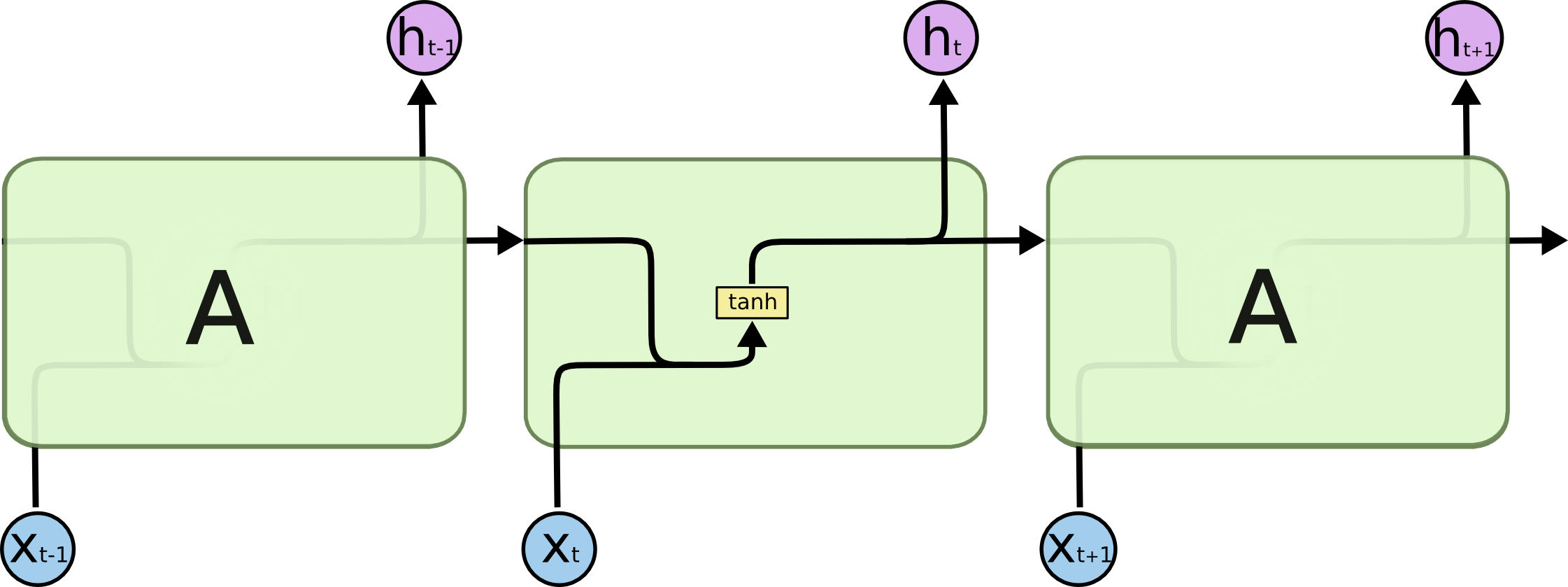
All the outputs before the final one are often discarded.
Say each prediction is a vector of size d, so \mathbf{y}_t \in \mathbb{R}^{1 \times d}.
Then the main equation of a SimpleRNN, given \mathbf{y}_0 = \mathbf{0}, is
\mathbf{y}_t = \psi\bigl( \mathbf{x}_t \mathbf{W}_x + \mathbf{y}_{t-1} \mathbf{W}_y + \mathbf{b} \bigr) .
Here, \begin{aligned} &\mathbf{x}_t \in \mathbb{R}^{1 \times m}, \mathbf{W}_x \in \mathbb{R}^{m \times d}, \\ &\mathbf{y}_{t-1} \in \mathbb{R}^{1 \times d}, \mathbf{W}_y \in \mathbb{R}^{d \times d}, \text{ and } \mathbf{b} \in \mathbb{R}^{d}. \end{aligned}
At each time step, a simple Recurrent Neural Network (RNN) takes an input vector x_t, incorporate it with the information from the previous hidden state {y}_{t-1} and produces an output vector at each time step y_t. The hidden state helps the network remember the context of the previous words, enabling it to make informed predictions about what comes next in the sequence. In a simple RNN, the output at time (t-1) is the same as the hidden state at time t.
The difference between RNN and RNNs with batch processing lies in the way how the neural network handles sequences of input data. With batch processing, the model processes multiple (b) input sequences simultaneously. The training data is grouped into batches, and the weights are updated based on the average error across the entire batch. Batch processing often results in more stable weight updates, as the model learns from a diverse set of examples in each batch, reducing the impact of noise in individual sequences.
Say we operate on batches of size b, then \mathbf{Y}_t \in \mathbb{R}^{b \times d}.
The main equation of a SimpleRNN, given \mathbf{Y}_0 = \mathbf{0}, is \mathbf{Y}_t = \psi\bigl( \mathbf{X}_t \mathbf{W}_x + \mathbf{Y}_{t-1} \mathbf{W}_y + \mathbf{b} \bigr) . Here, \begin{aligned} &\mathbf{X}_t \in \mathbb{R}^{b \times m}, \mathbf{W}_x \in \mathbb{R}^{m \times d}, \\ &\mathbf{Y}_{t-1} \in \mathbb{R}^{b \times d}, \mathbf{W}_y \in \mathbb{R}^{d \times d}, \text{ and } \mathbf{b} \in \mathbb{R}^{d}. \end{aligned}
1num_obs = 4
2num_time_steps = 3
3num_time_series = 2
X = np.arange(num_obs*num_time_steps*num_time_series).astype(np.float32) \
4 .reshape([num_obs, num_time_steps, num_time_series])
output_size = 1
y = np.array([0, 0, 1, 1])1X[:2]X[:2] selects the first two slices (0 and 1) along the first dimension, and returns a sub-tensor of shape (2,3,2).
array([[[ 0., 1.],
[ 2., 3.],
[ 4., 5.]],
[[ 6., 7.],
[ 8., 9.],
[10., 11.]]], dtype=float32)1X[2:]X[2:] selects the last two slices (2 and 3) along the first dimension, and returns a sub-tensor of shape (2,3,2).
array([[[12., 13.],
[14., 15.],
[16., 17.]],
[[18., 19.],
[20., 21.],
[22., 23.]]], dtype=float32)As usual, the SimpleRNN is just a layer in Keras.
1from keras.layers import SimpleRNN
2random.seed(1234)
model = Sequential([
3 SimpleRNN(output_size, activation="sigmoid")
])
4model.compile(loss="binary_crossentropy", metrics=["accuracy"])
5hist = model.fit(X, y, epochs=500, verbose=False)
6model.evaluate(X, y, verbose=False)hist
[8.05906867980957, 0.5]The predicted probabilities on the training set are:
model.predict(X, verbose=0)array([[8.56e-05],
[2.25e-10],
[5.98e-16],
[1.59e-21]], dtype=float32)To verify the results of predicted probabilities, we can obtain the weights of the fitted model and calculate the outcome manually as follows.
model.get_weights()[array([[-1.47],
[-0.67]], dtype=float32),
array([[0.99]], dtype=float32),
array([-0.14], dtype=float32)]def sigmoid(x):
return 1 / (1 + np.exp(-x))
W_x, W_y, b = model.get_weights()
Y = np.zeros((num_obs, output_size), dtype=np.float32)
for t in range(num_time_steps):
X_t = X[:, t, :]
z = X_t @ W_x + Y @ W_y + b
Y = sigmoid(z)
Yarray([[8.56e-05],
[2.25e-10],
[5.98e-16],
[1.59e-21]], dtype=float32)Simple RNN structures encounter vanishing gradient problems, hence, struggle with learning long term dependencies. LSTM are designed to overcome this problem. LSTMs have a more complex network structure (contains more memory cells and gating mechanisms) and can better regulate the information flow.
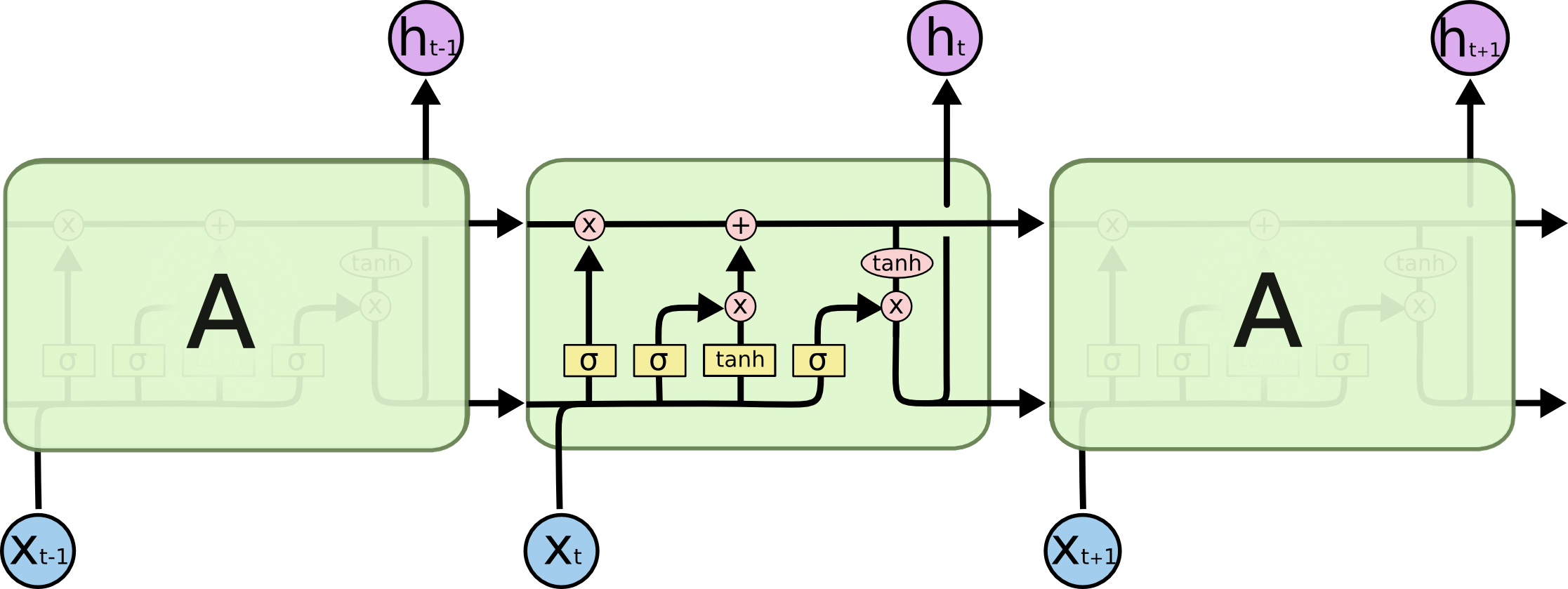
GRUs are simpler compared to LSTM, hence, computationally more efficient than LSTMs.

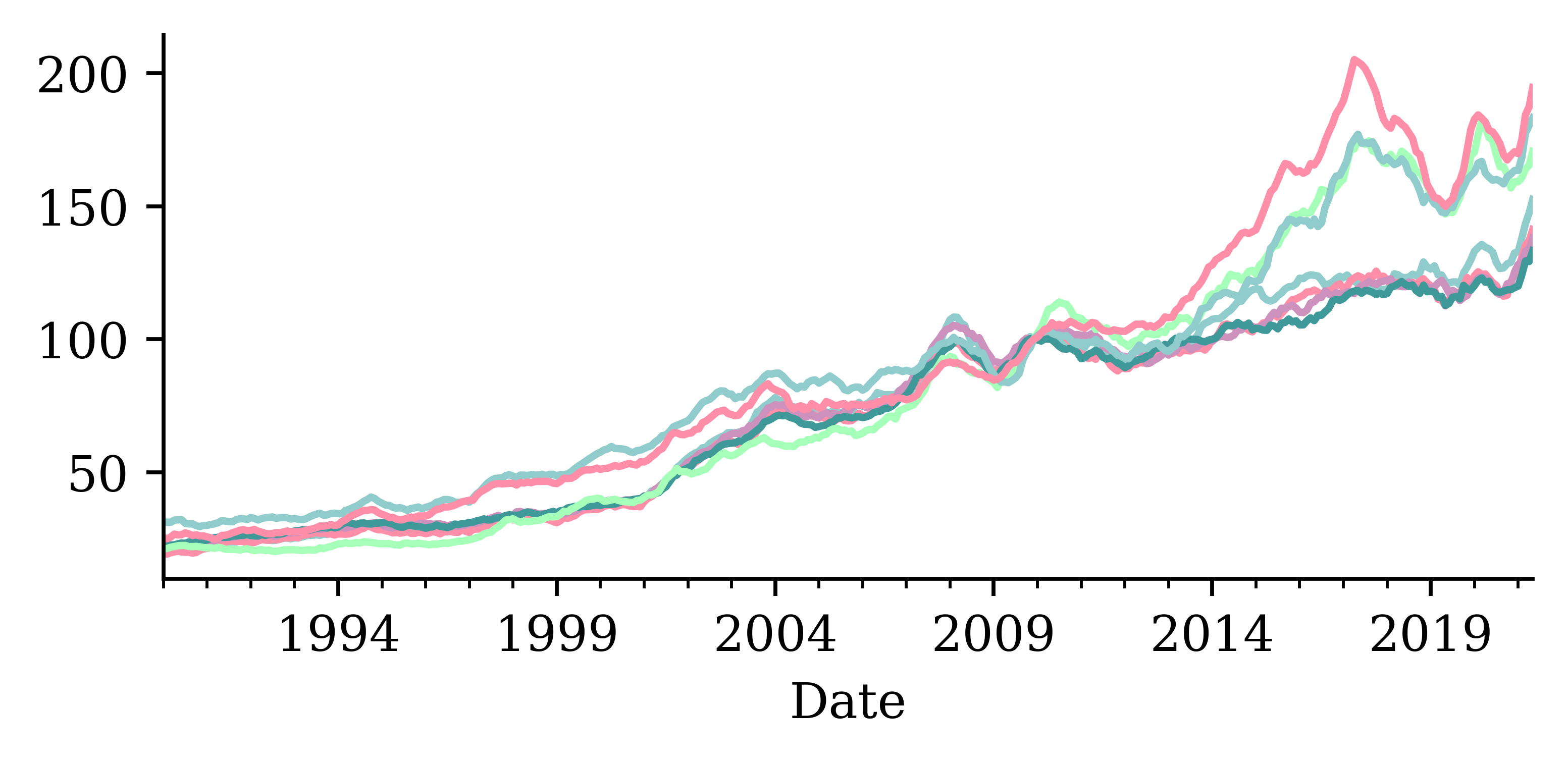
changes = house_prices.pct_change().dropna()
changes.round(2)| Brisbane | East_Bris | North_Bris | West_Bris | Melbourne | North_Syd | Sydney | |
|---|---|---|---|---|---|---|---|
| Date | |||||||
| 1990-02-28 | 0.03 | -0.01 | 0.01 | 0.01 | 0.00 | -0.00 | -0.02 |
| 1990-03-31 | 0.01 | 0.03 | 0.01 | 0.01 | 0.02 | -0.00 | 0.03 |
| ... | ... | ... | ... | ... | ... | ... | ... |
| 2021-04-30 | 0.03 | 0.01 | 0.01 | -0.00 | 0.01 | 0.02 | 0.02 |
| 2021-05-31 | 0.03 | 0.03 | 0.03 | 0.03 | 0.03 | 0.02 | 0.04 |
376 rows × 7 columns
changes.plot();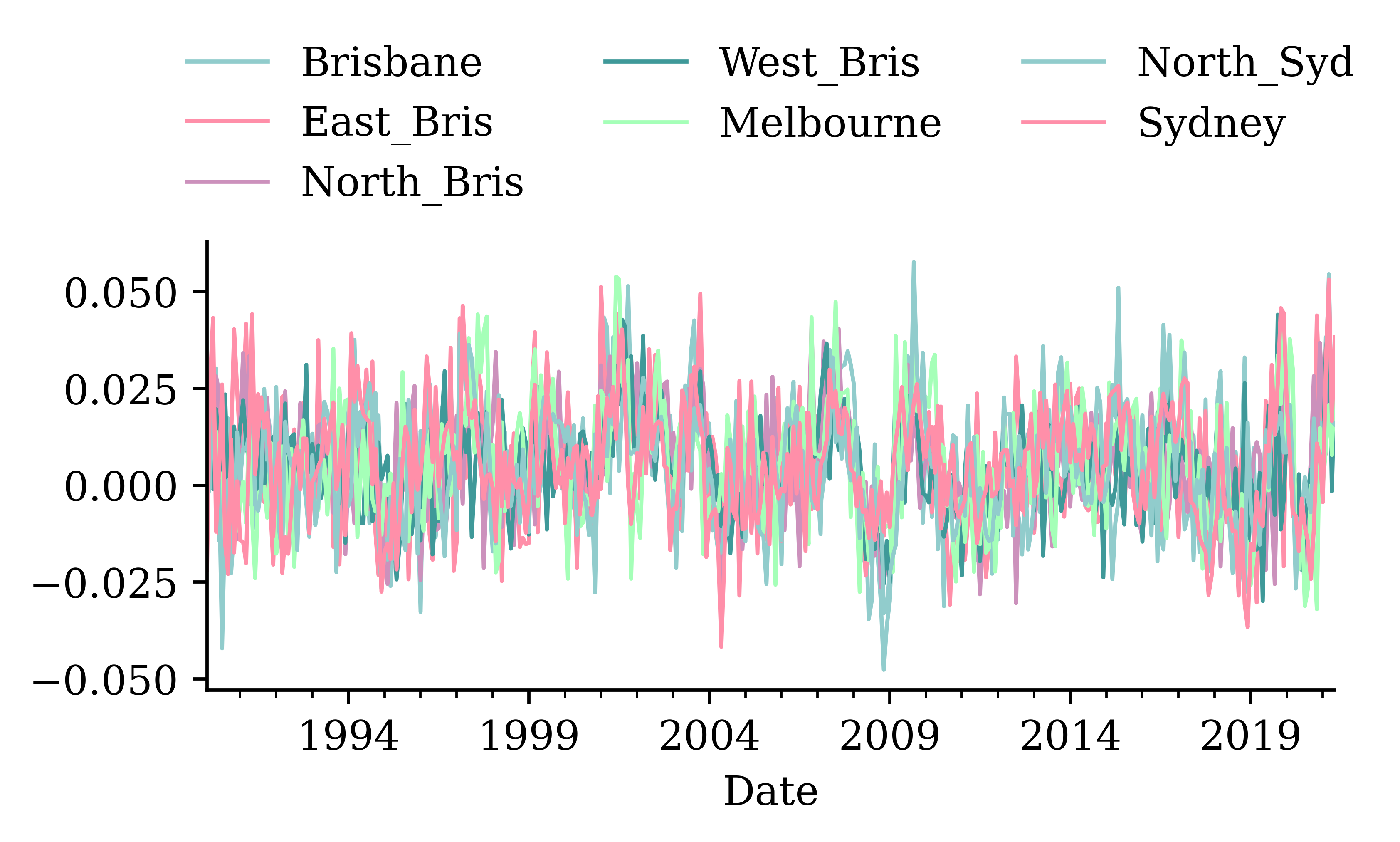
changes.mean()Brisbane 0.005496
East_Bris 0.005416
North_Bris 0.005024
West_Bris 0.004842
Melbourne 0.005677
North_Syd 0.004819
Sydney 0.005526
dtype: float64changes *= 100changes.mean()Brisbane 0.549605
East_Bris 0.541562
North_Bris 0.502390
West_Bris 0.484204
Melbourne 0.567700
North_Syd 0.481863
Sydney 0.552641
dtype: float64changes.plot(legend=False);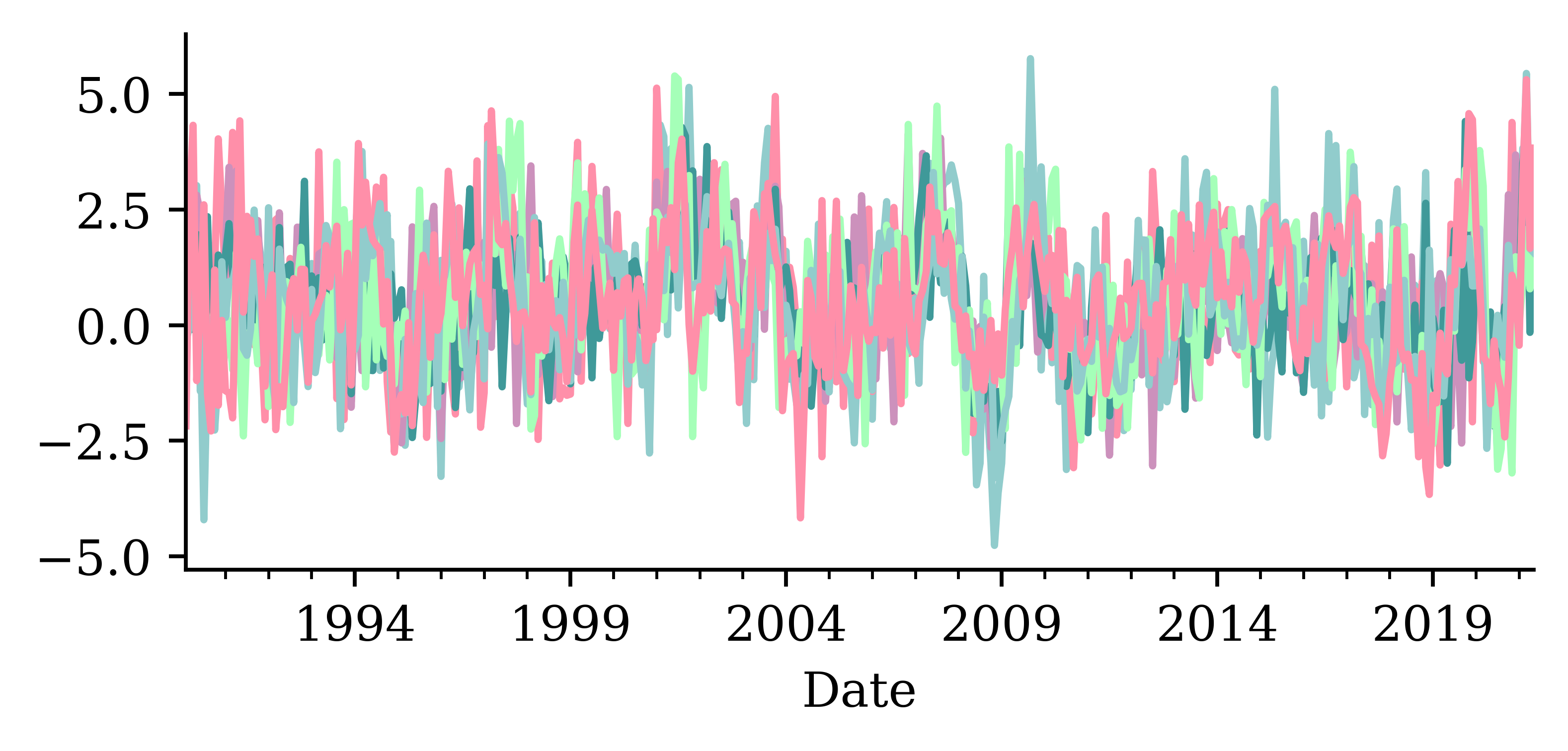
num_train = int(0.6 * len(changes))
num_val = int(0.2 * len(changes))
num_test = len(changes) - num_train - num_val
print(f"# Train: {num_train}, # Val: {num_val}, # Test: {num_test}")# Train: 225, # Val: 75, # Test: 76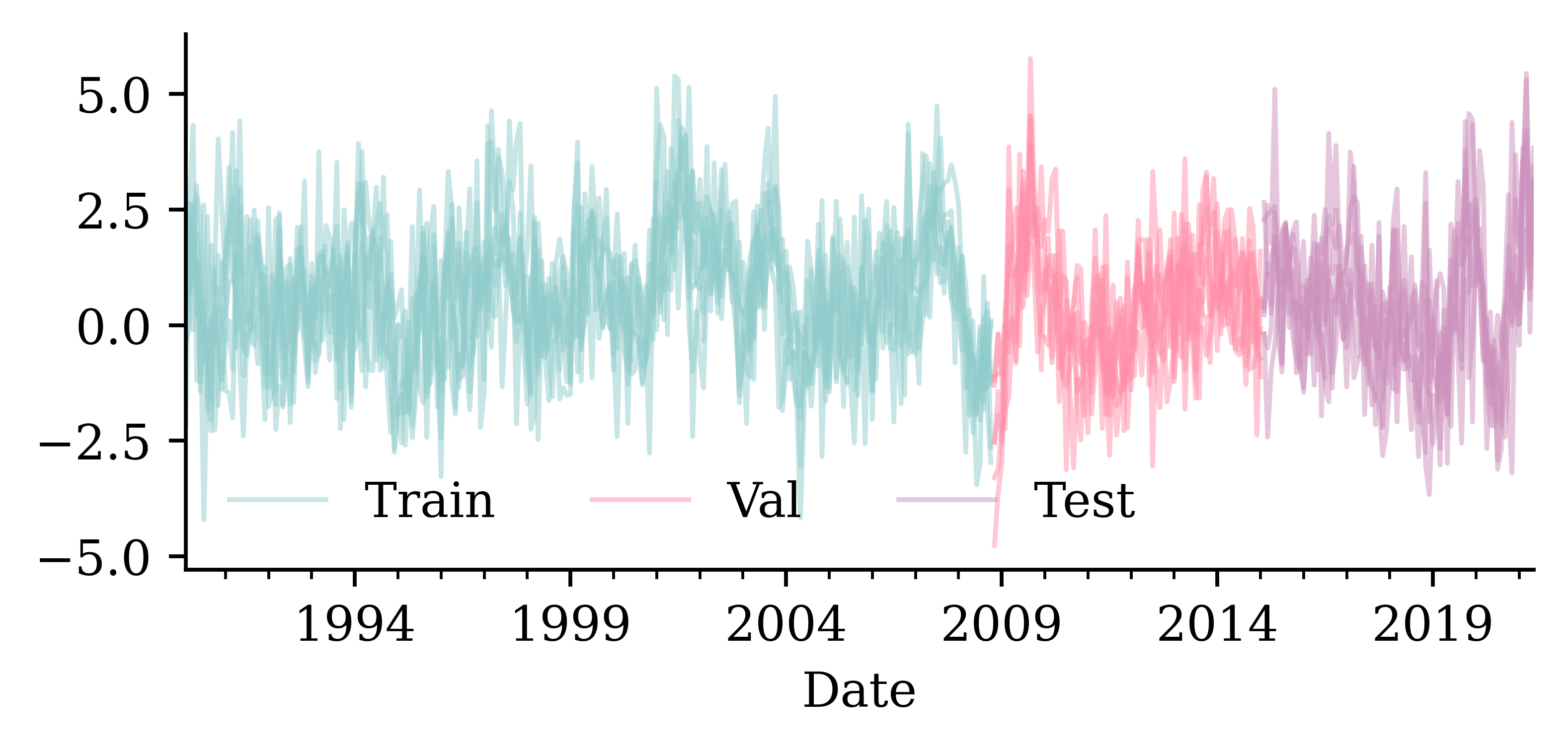
Keras has a built-in method for converting a time series into subsequences/chunks.
from keras.utils import timeseries_dataset_from_array
integers = range(10)
dummy_dataset = timeseries_dataset_from_array(
data=integers[:-3],
targets=integers[3:],
sequence_length=3,
batch_size=2,
)
for inputs, targets in dummy_dataset:
for i in range(inputs.shape[0]):
print([int(x) for x in inputs[i]], int(targets[i]))[0, 1, 2] 3
[1, 2, 3] 4
[2, 3, 4] 5
[3, 4, 5] 6
[4, 5, 6] 7If you have a lot of time series data, then use:
from keras.utils import timeseries_dataset_from_array
data = range(20); seq = 3; ts = data[:-seq]; target = data[seq:]
nTrain = int(0.5 * len(ts)); nVal = int(0.25 * len(ts))
nTest = len(ts) - nTrain - nVal
print(f"# Train: {nTrain}, # Val: {nVal}, # Test: {nTest}")# Train: 8, # Val: 4, # Test: 5trainDS = \
timeseries_dataset_from_array(
ts, target, seq,
end_index=nTrain)valDS = \
timeseries_dataset_from_array(
ts, target, seq,
start_index=nTrain,
end_index=nTrain+nVal)testDS = \
timeseries_dataset_from_array(
ts, target, seq,
start_index=nTrain+nVal)Training dataset
[0, 1, 2] 3
[1, 2, 3] 4
[2, 3, 4] 5
[3, 4, 5] 6
[4, 5, 6] 7
[5, 6, 7] 8Validation dataset
[8, 9, 10] 11
[9, 10, 11] 12Test dataset
[12, 13, 14] 15
[13, 14, 15] 16
[14, 15, 16] 17If you don’t have a lot of time series data, consider:
X = []; y = []
for i in range(len(data)-seq):
X.append(data[i:i+seq])
y.append(data[i+seq])
X = np.array(X); y = np.array(y);nTrain = int(0.5 * X.shape[0])
X_train = X[:nTrain]
y_train = y[:nTrain]nVal = int(np.ceil(0.25 * X.shape[0]))
X_val = X[nTrain:nTrain+nVal]
y_val = y[nTrain:nTrain+nVal]nTest = X.shape[0] - nTrain - nVal
X_test = X[nTrain+nVal:]
y_test = y[nTrain+nVal:]Training dataset
[0, 1, 2] 3
[1, 2, 3] 4
[2, 3, 4] 5
[3, 4, 5] 6
[4, 5, 6] 7
[5, 6, 7] 8
[6, 7, 8] 9
[7, 8, 9] 10Validation dataset
[8, 9, 10] 11
[9, 10, 11] 12
[10, 11, 12] 13
[11, 12, 13] 14
[12, 13, 14] 15Test dataset
[13, 14, 15] 16
[14, 15, 16] 17
[15, 16, 17] 18
[16, 17, 18] 19# Num. of input time series.
num_ts = changes.shape[1]
# How many prev. months to use.
seq_length = 6
# Predict the next month ahead.
ahead = 1
# The index of the first target.
delay = (seq_length+ahead-1)# Which suburb to predict.
target_suburb = changes["Sydney"]
train_ds = \
timeseries_dataset_from_array(
changes[:-delay],
targets=target_suburb[delay:],
sequence_length=seq_length,
end_index=num_train)val_ds = \
timeseries_dataset_from_array(
changes[:-delay],
targets=target_suburb[delay:],
sequence_length=seq_length,
start_index=num_train,
end_index=num_train+num_val)test_ds = \
timeseries_dataset_from_array(
changes[:-delay],
targets=target_suburb[delay:],
sequence_length=seq_length,
start_index=num_train+num_val)Dataset to numpyThe Dataset object can be handed to Keras directly, but if we really need a numpy array, we can run:
X_train = np.concatenate(list(train_ds.map(lambda x, y: x)))
y_train = np.concatenate(list(train_ds.map(lambda x, y: y)))The shape of our training set is now:
X_train.shape(220, 6, 7)y_train.shape(220,)Later, we need the targets as numpy arrays:
y_train = np.concatenate(list(train_ds.map(lambda x, y: y)))
y_val = np.concatenate(list(val_ds.map(lambda x, y: y)))
y_test = np.concatenate(list(test_ds.map(lambda x, y: y)))from keras.layers import Input, Flatten
random.seed(1)
model_dense = Sequential([
Input((seq_length, num_ts)),
Flatten(),
Dense(50, activation="leaky_relu"),
Dense(20, activation="leaky_relu"),
Dense(1, activation="linear")
])
model_dense.compile(loss="mse", optimizer="adam")
print(f"This model has {model_dense.count_params()} parameters.")
es = EarlyStopping(patience=50, restore_best_weights=True, verbose=1)
%time hist = model_dense.fit(train_ds, epochs=1_000, \
validation_data=val_ds, callbacks=[es], verbose=0);This model has 3191 parameters.
Epoch 126: early stopping
Restoring model weights from the end of the best epoch: 76.
CPU times: user 6.48 s, sys: 3.55 s, total: 10 s
Wall time: 3.4 sfrom keras.utils import plot_model
plot_model(model_dense, show_shapes=True)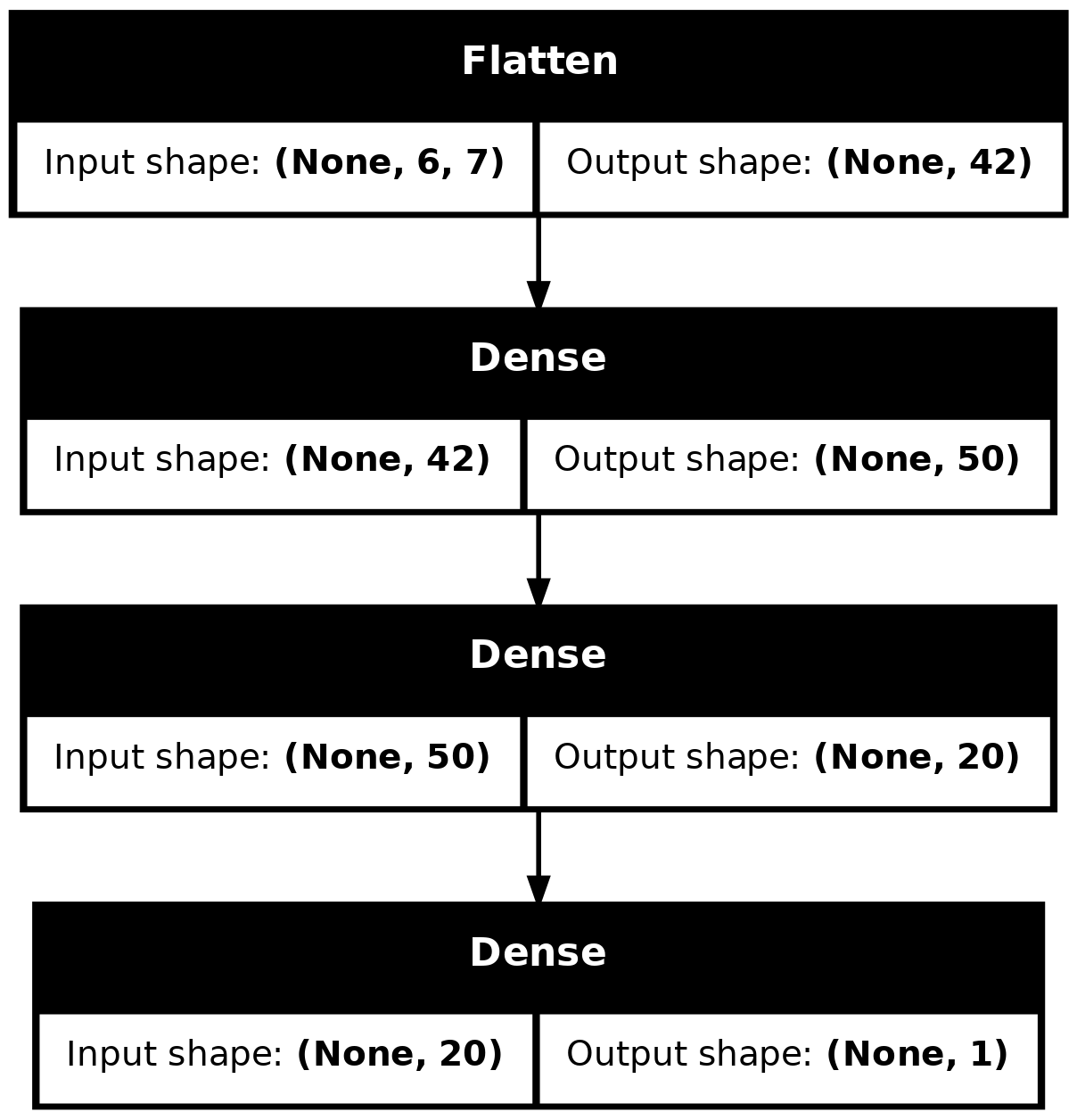
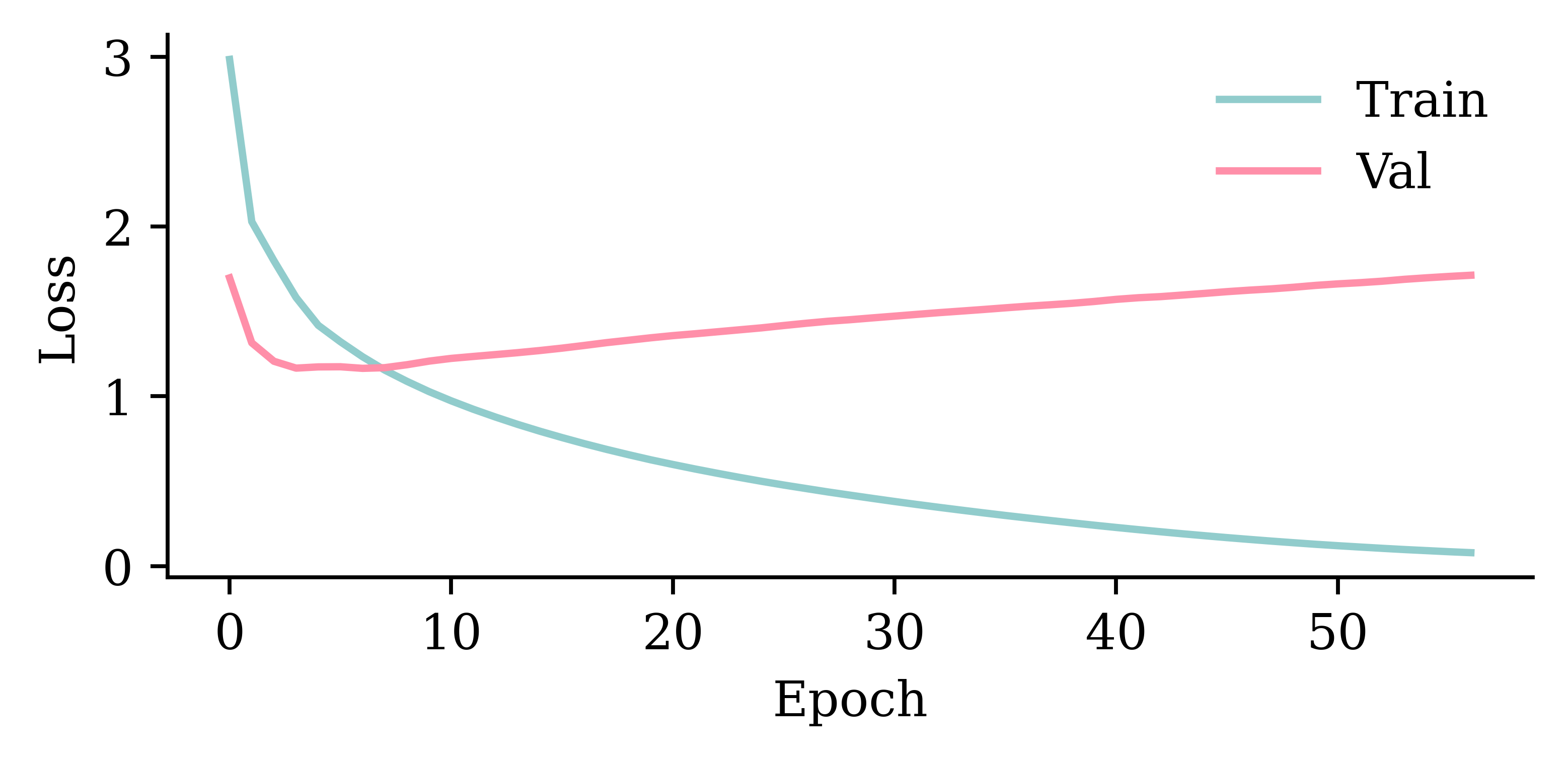
model_dense.evaluate(val_ds, verbose=0)1.3521653413772583
SimpleRNN layerrandom.seed(1)
model_simple = Sequential([
Input((seq_length, num_ts)),
SimpleRNN(50),
Dense(1, activation="linear")
])
model_simple.compile(loss="mse", optimizer="adam")
print(f"This model has {model_simple.count_params()} parameters.")
es = EarlyStopping(patience=50, restore_best_weights=True, verbose=1)
%time hist = model_simple.fit(train_ds, epochs=1_000, \
validation_data=val_ds, callbacks=[es], verbose=0);This model has 2951 parameters.
Epoch 57: early stopping
Restoring model weights from the end of the best epoch: 7.
CPU times: user 3.07 s, sys: 1.54 s, total: 4.61 s
Wall time: 1.71 s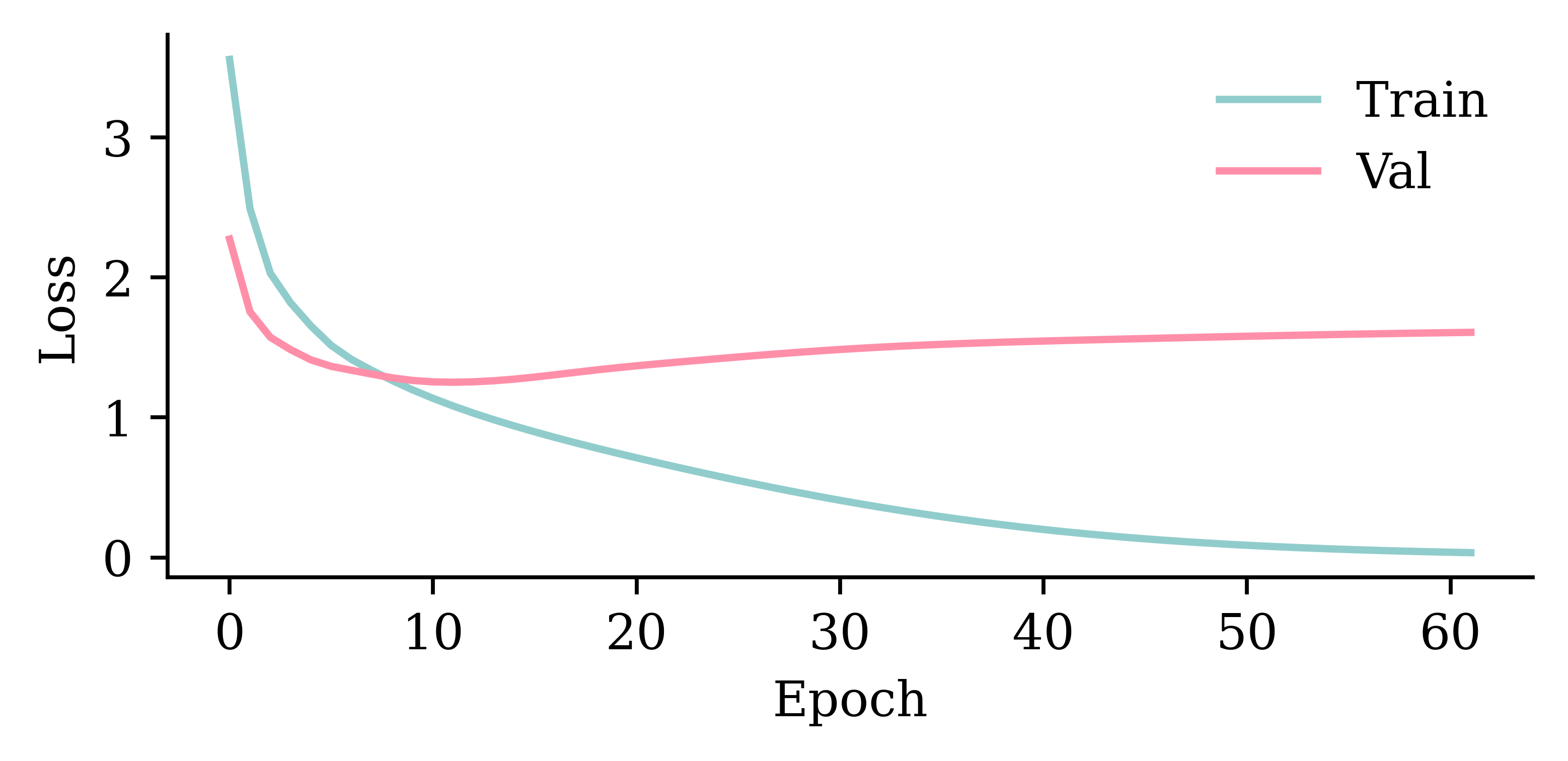
model_simple.evaluate(val_ds, verbose=0)0.8697079420089722plot_model(model_simple, show_shapes=True)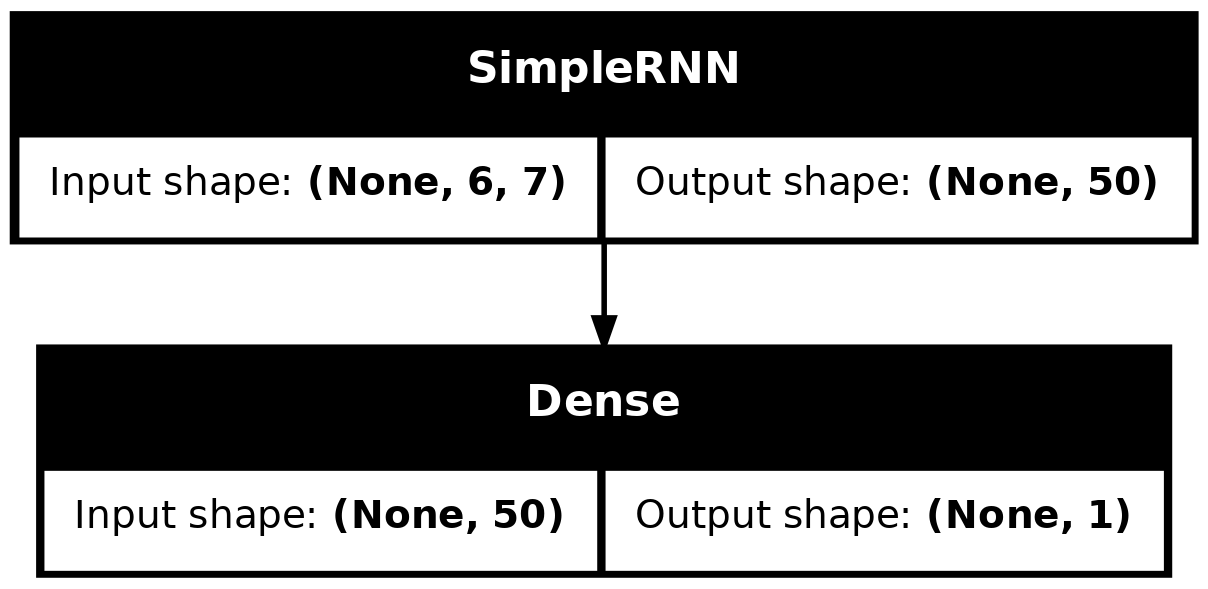
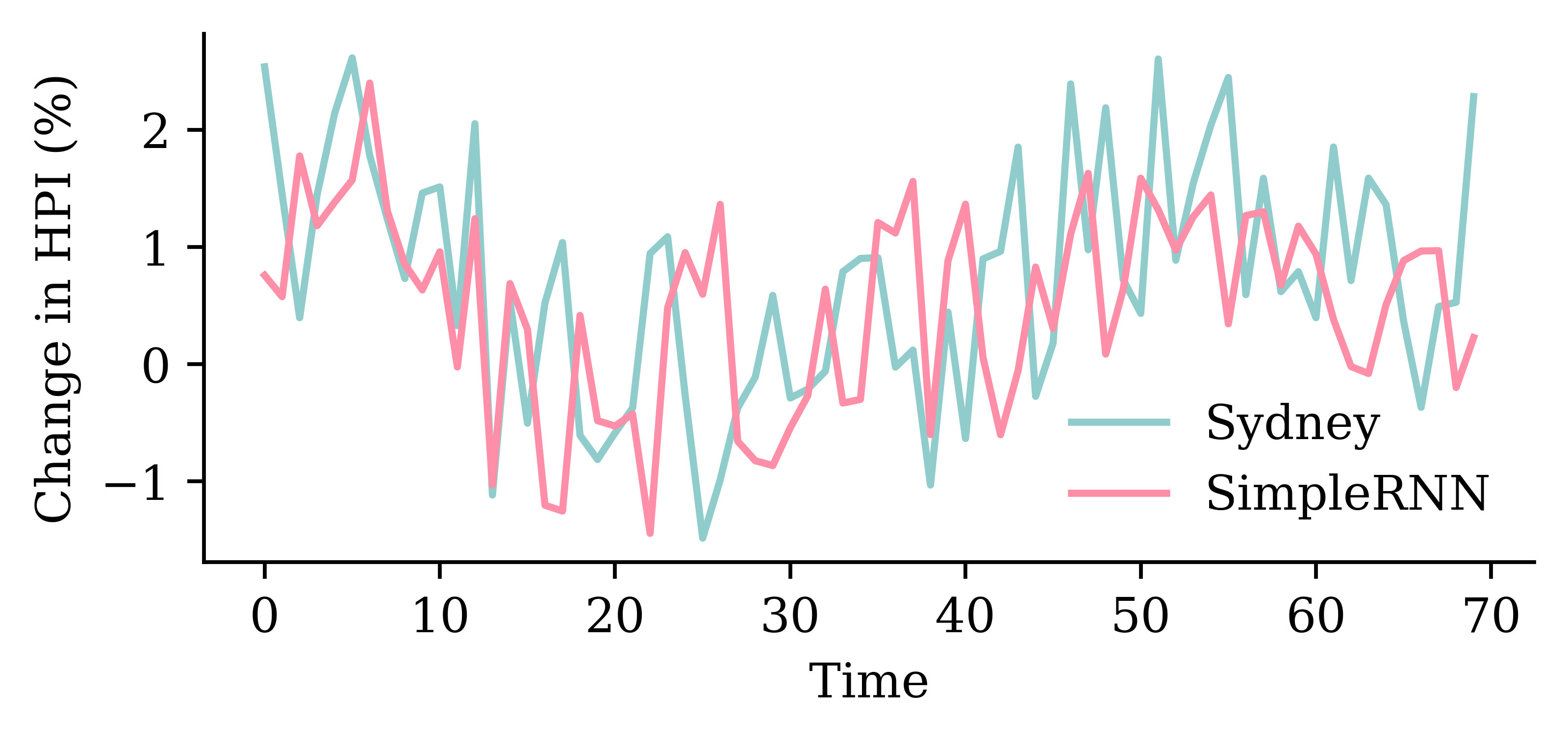
LSTM layerfrom keras.layers import LSTM
random.seed(1)
model_lstm = Sequential([
Input((seq_length, num_ts)),
LSTM(50),
Dense(1, activation="linear")
])
model_lstm.compile(loss="mse", optimizer="adam")
es = EarlyStopping(patience=50, restore_best_weights=True, verbose=1)
%time hist = model_lstm.fit(train_ds, epochs=1_000, \
validation_data=val_ds, callbacks=[es], verbose=0);Epoch 72: early stopping
Restoring model weights from the end of the best epoch: 22.
CPU times: user 4.09 s, sys: 2 s, total: 6.09 s
Wall time: 2.39 s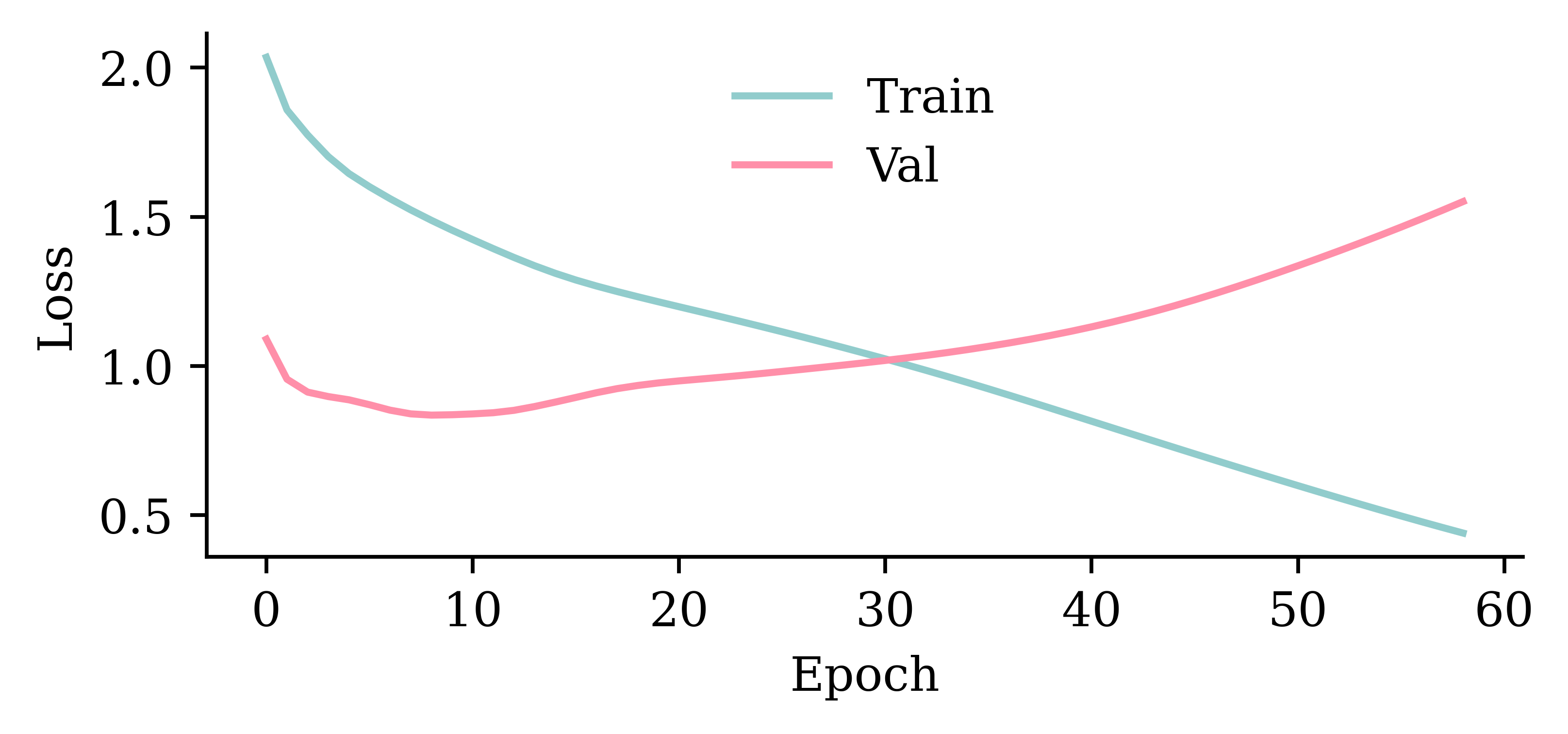
model_lstm.evaluate(val_ds, verbose=0)0.8331283330917358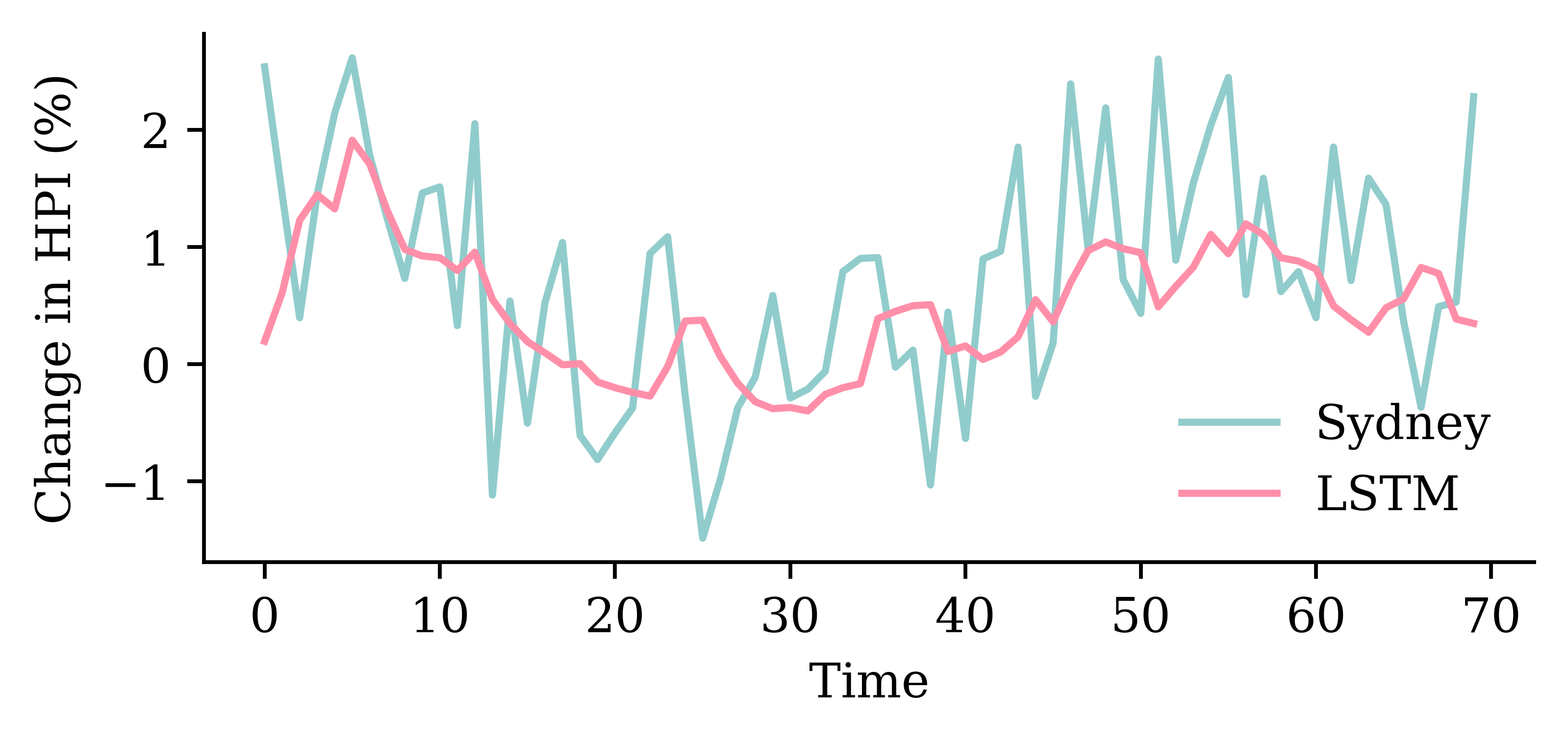
GRU layerfrom keras.layers import GRU
random.seed(1)
model_gru = Sequential([
Input((seq_length, num_ts)),
GRU(50),
Dense(1, activation="linear")
])
model_gru.compile(loss="mse", optimizer="adam")
es = EarlyStopping(patience=50, restore_best_weights=True, verbose=1)
%time hist = model_gru.fit(train_ds, epochs=1_000, \
validation_data=val_ds, callbacks=[es], verbose=0)Epoch 66: early stopping
Restoring model weights from the end of the best epoch: 16.
CPU times: user 3.81 s, sys: 1.82 s, total: 5.63 s
Wall time: 2.21 s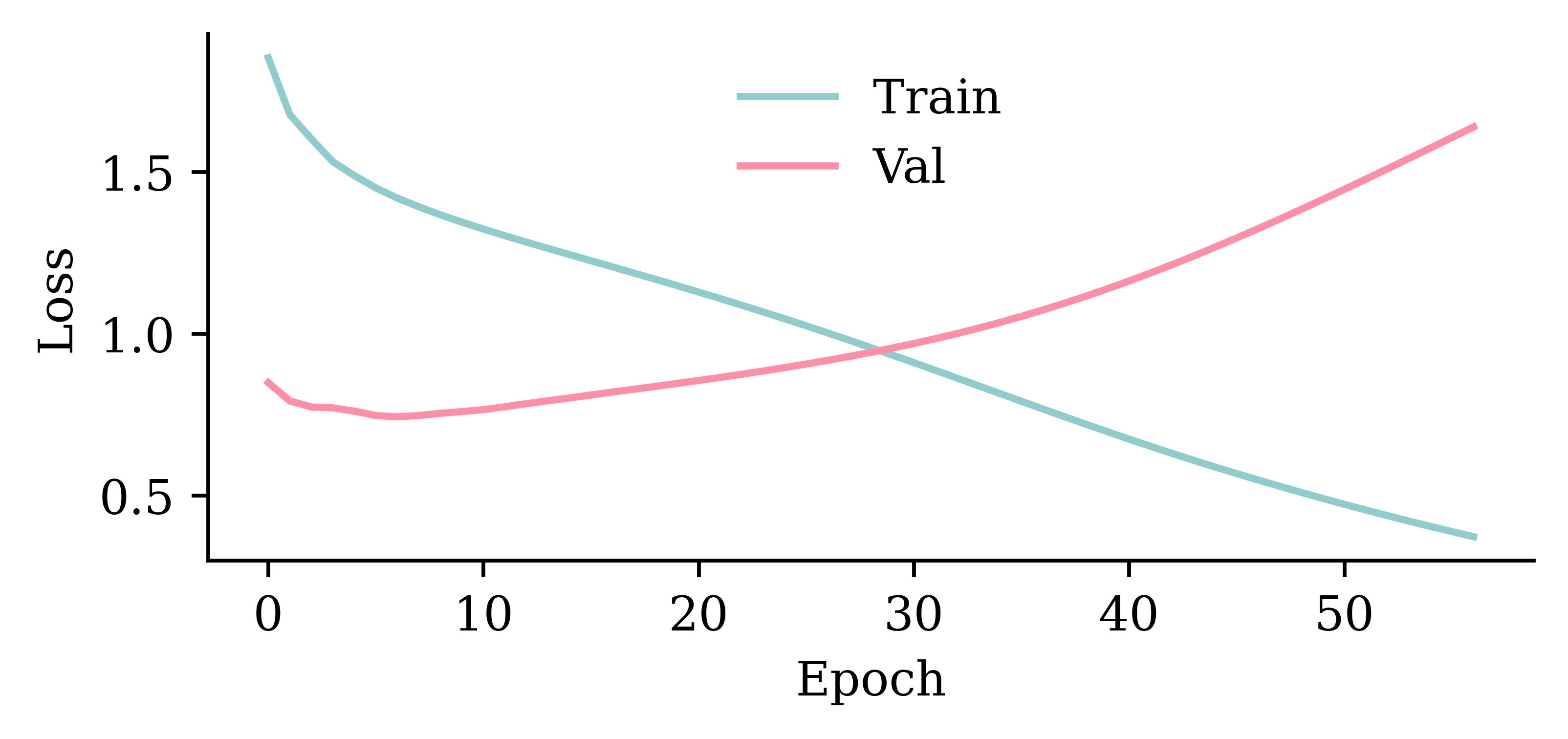
model_gru.evaluate(val_ds, verbose=0)0.8188608884811401
GRU layersrandom.seed(1)
model_two_grus = Sequential([
Input((seq_length, num_ts)),
GRU(50, return_sequences=True),
GRU(50),
Dense(1, activation="linear")
])
model_two_grus.compile(loss="mse", optimizer="adam")
es = EarlyStopping(patience=50, restore_best_weights=True, verbose=1)
%time hist = model_two_grus.fit(train_ds, epochs=1_000, \
validation_data=val_ds, callbacks=[es], verbose=0)Epoch 69: early stopping
Restoring model weights from the end of the best epoch: 19.
CPU times: user 4.5 s, sys: 1.93 s, total: 6.43 s
Wall time: 2.88 s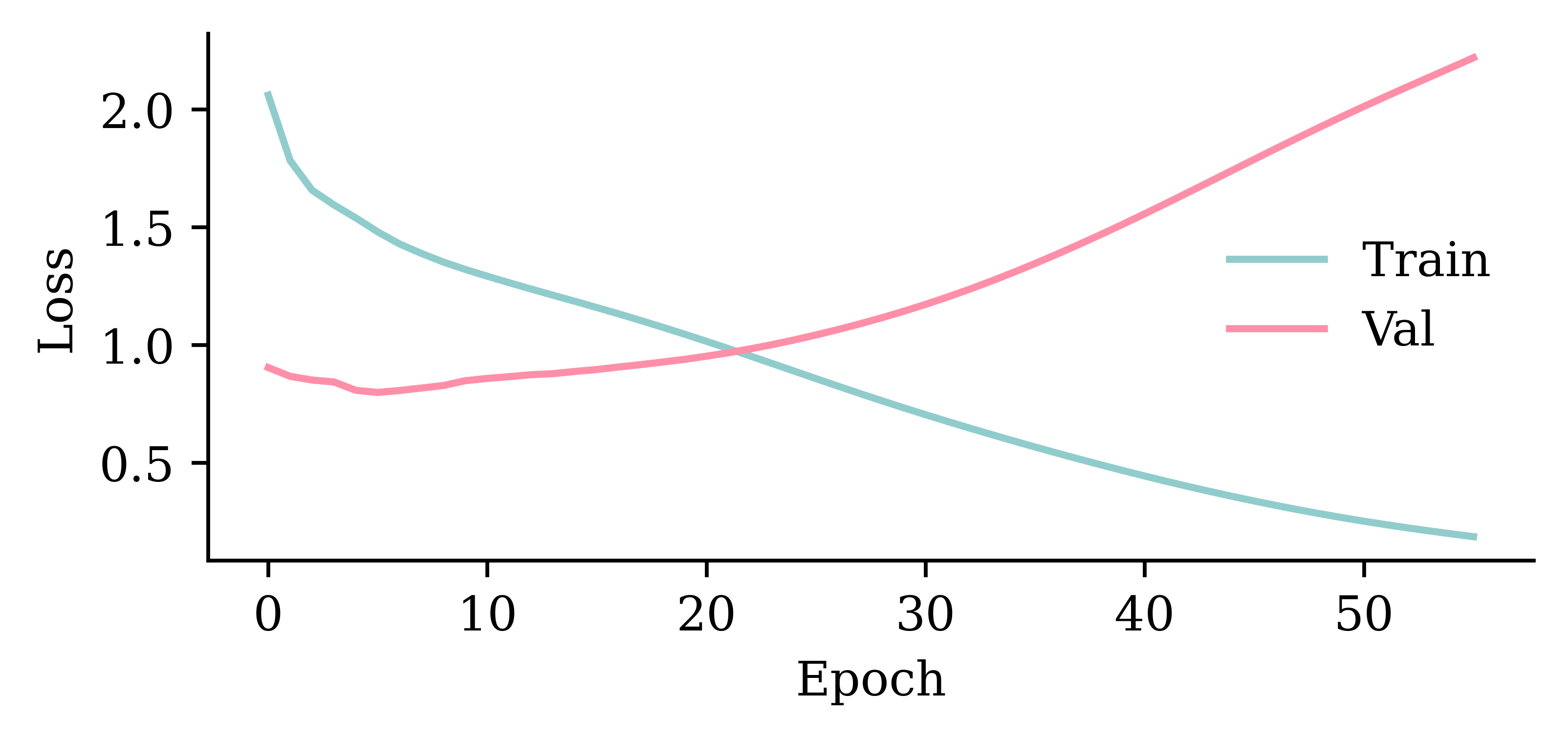
model_two_grus.evaluate(val_ds, verbose=0)0.777770459651947
| Model | MSE | |
|---|---|---|
| 0 | Dense | 1.352165 |
| 1 | SimpleRNN | 0.869708 |
| 2 | LSTM | 0.833128 |
| 3 | GRU | 0.818861 |
| 4 | 2 GRUs | 0.777770 |
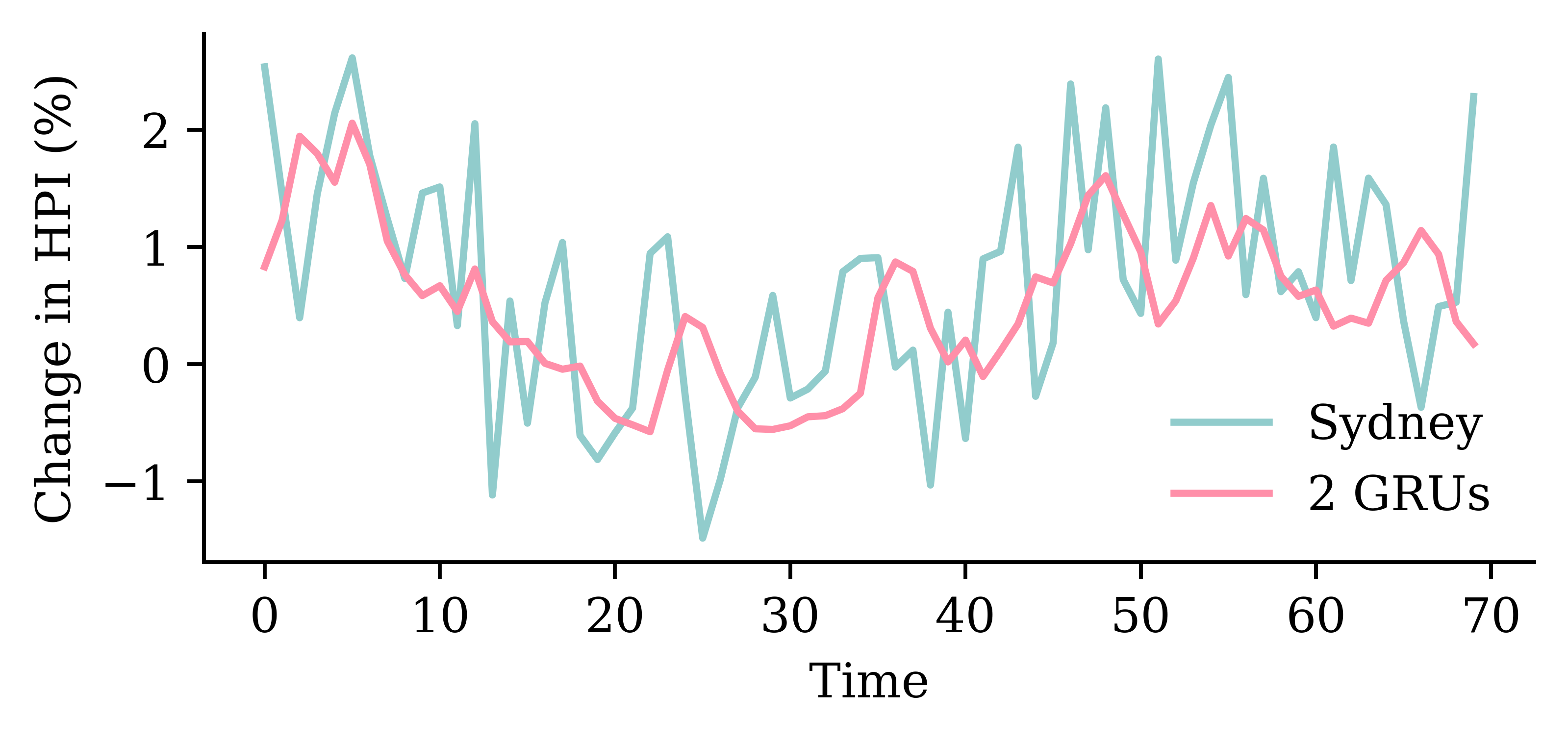
The network with two GRU layers is the best.
model_two_grus.evaluate(test_ds, verbose=0)1.9774343967437744
Change the targets argument to include all the suburbs.
train_ds = \
timeseries_dataset_from_array(
changes[:-delay],
targets=changes[delay:],
sequence_length=seq_length,
end_index=num_train)val_ds = \
timeseries_dataset_from_array(
changes[:-delay],
targets=changes[delay:],
sequence_length=seq_length,
start_index=num_train,
end_index=num_train+num_val)test_ds = \
timeseries_dataset_from_array(
changes[:-delay],
targets=changes[delay:],
sequence_length=seq_length,
start_index=num_train+num_val)Dataset to numpyThe shape of our training set is now:
X_train = np.concatenate(list(train_ds.map(lambda x, y: x)))
X_train.shape(220, 6, 7)Y_train = np.concatenate(list(train_ds.map(lambda x, y: y)))
Y_train.shape(220, 7)Later, we need the targets as numpy arrays:
Y_train = np.concatenate(list(train_ds.map(lambda x, y: y)))
Y_val = np.concatenate(list(val_ds.map(lambda x, y: y)))
Y_test = np.concatenate(list(test_ds.map(lambda x, y: y)))random.seed(1)
model_dense = Sequential([
Input((seq_length, num_ts)),
Flatten(),
Dense(50, activation="leaky_relu"),
Dense(20, activation="leaky_relu"),
Dense(num_ts, activation="linear")
])
model_dense.compile(loss="mse", optimizer="adam")
print(f"This model has {model_dense.count_params()} parameters.")
es = EarlyStopping(patience=50, restore_best_weights=True, verbose=1)
%time hist = model_dense.fit(train_ds, epochs=1_000, \
validation_data=val_ds, callbacks=[es], verbose=0);This model has 3317 parameters.
Epoch 96: early stopping
Restoring model weights from the end of the best epoch: 46.
CPU times: user 4.97 s, sys: 2.69 s, total: 7.66 s
Wall time: 2.65 splot_model(model_dense, show_shapes=True)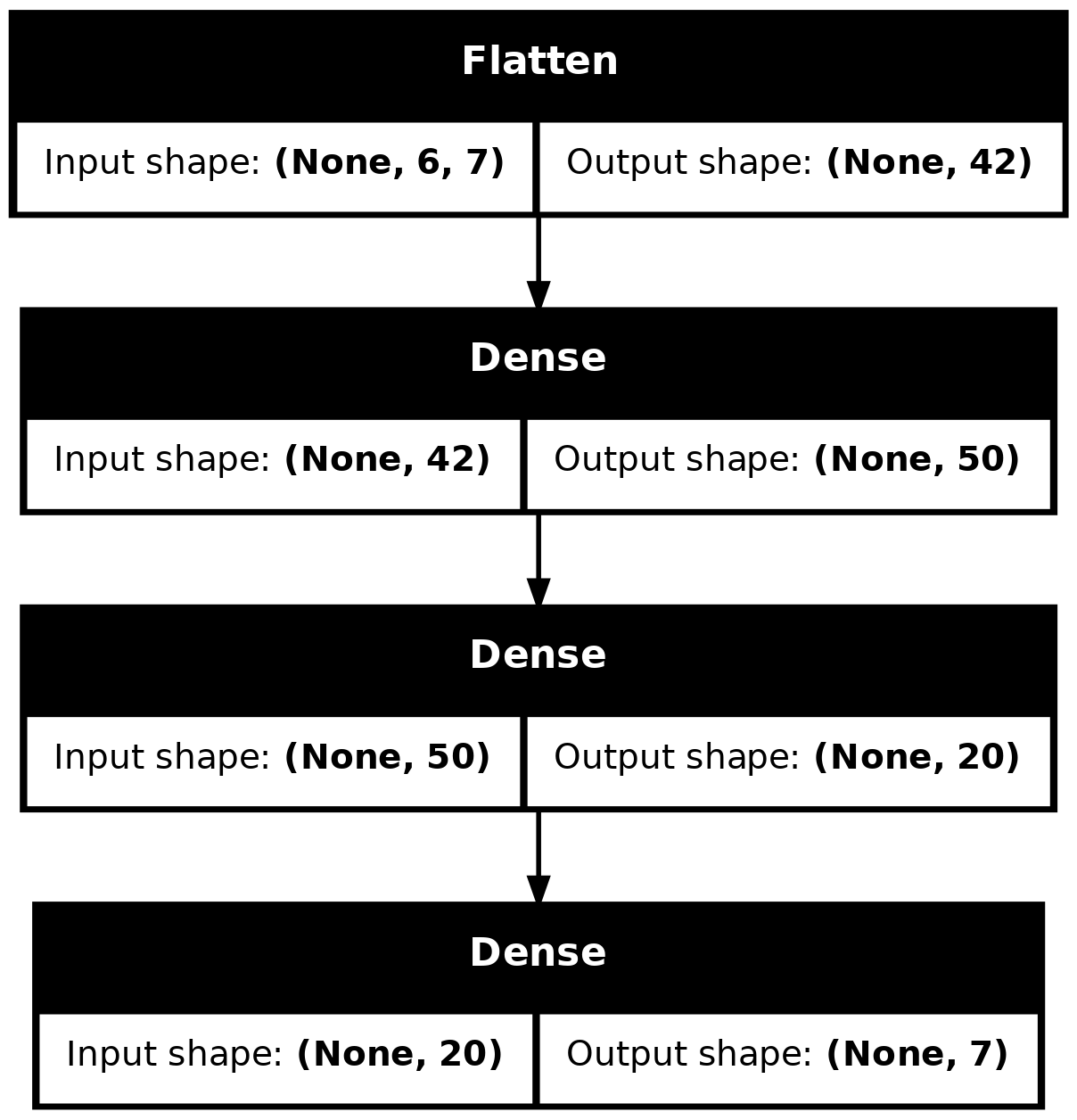
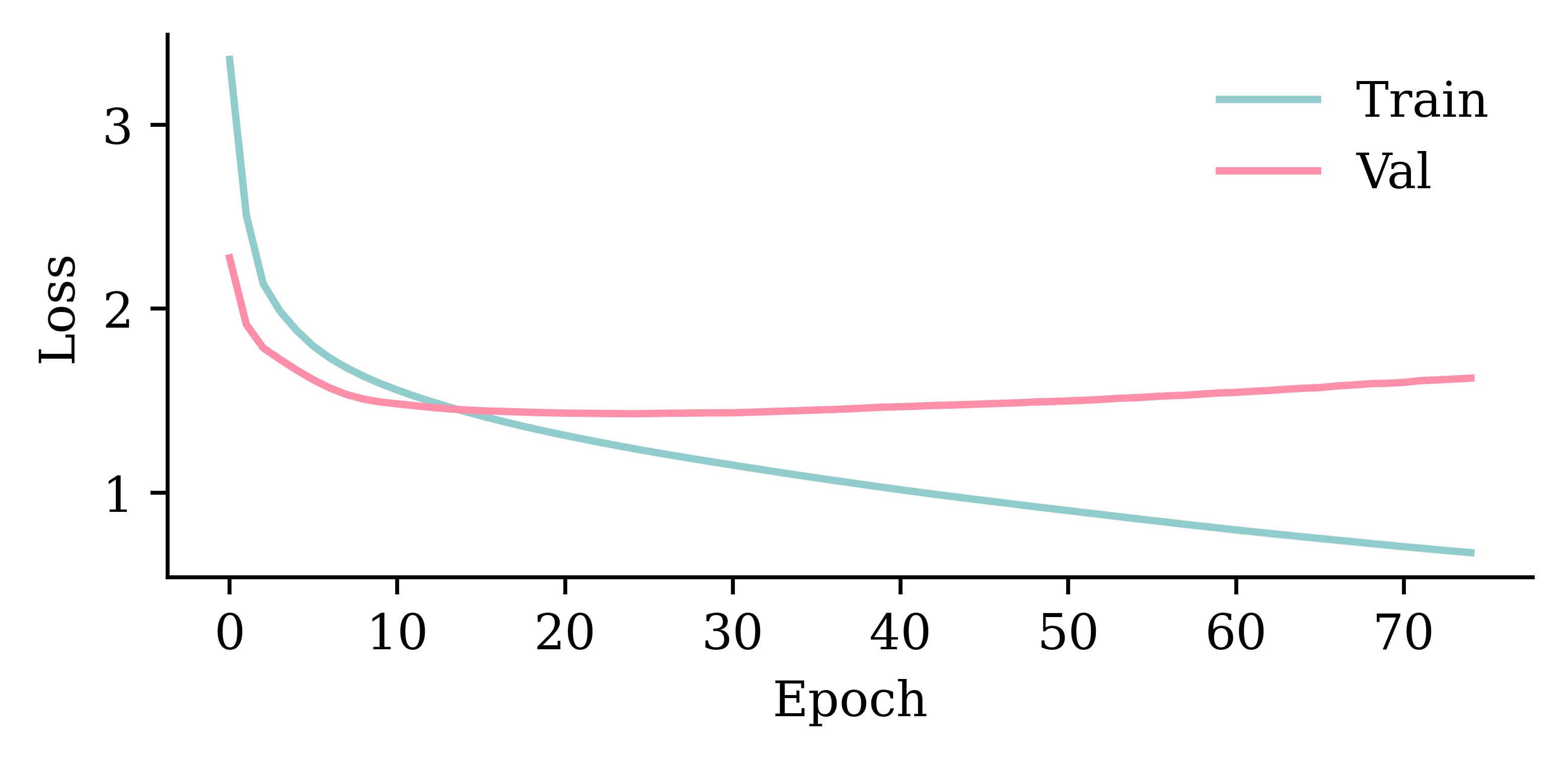
model_dense.evaluate(val_ds, verbose=0)1.445832371711731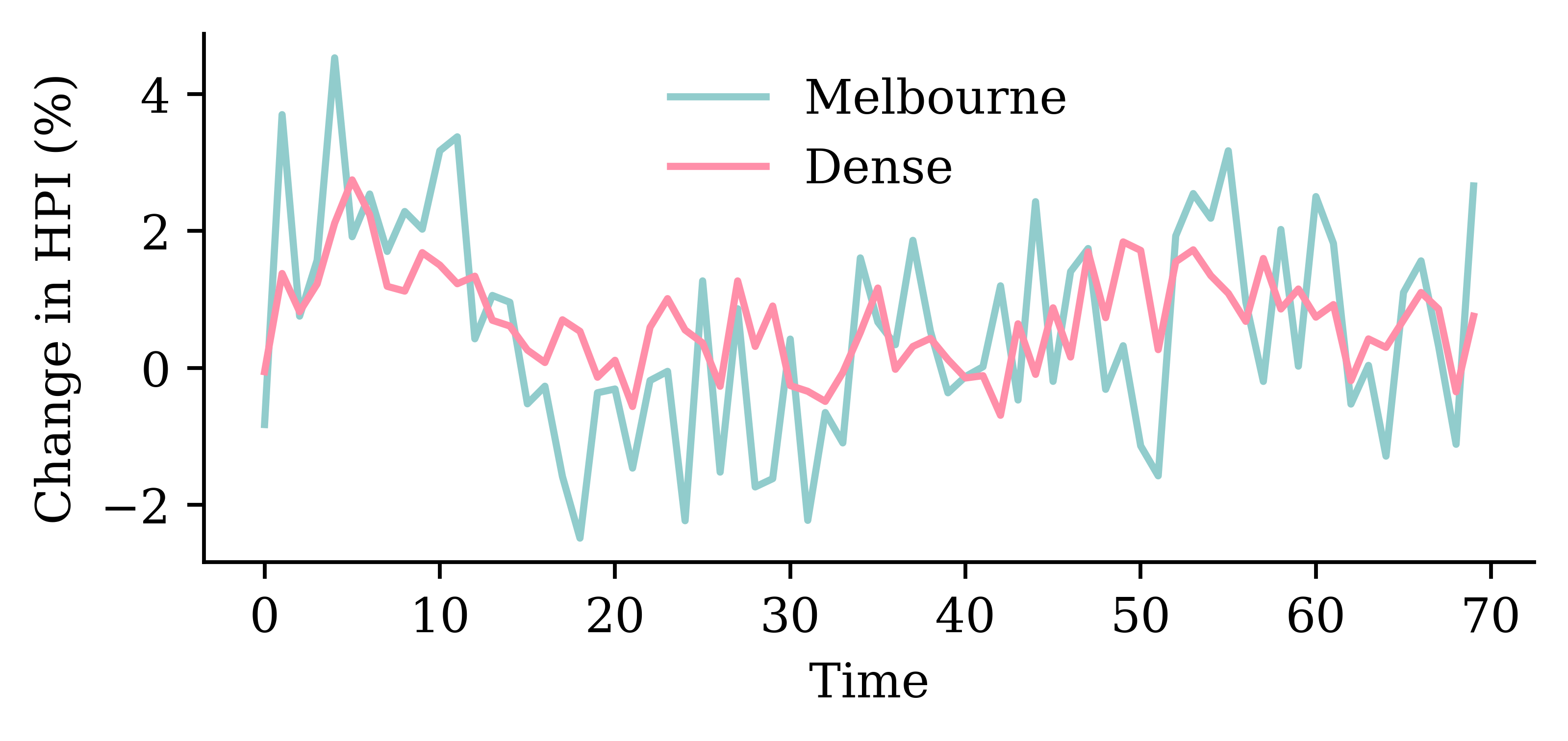

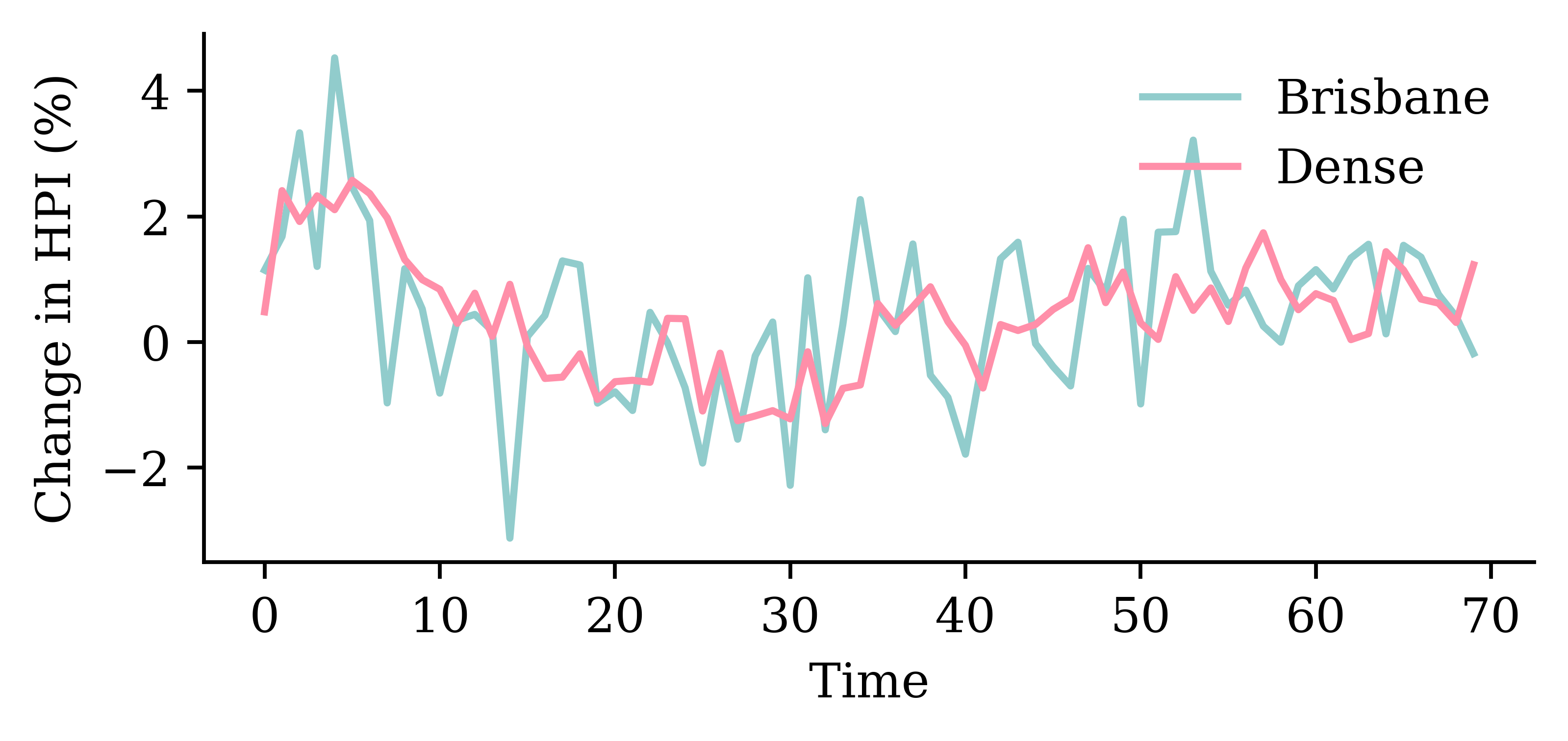
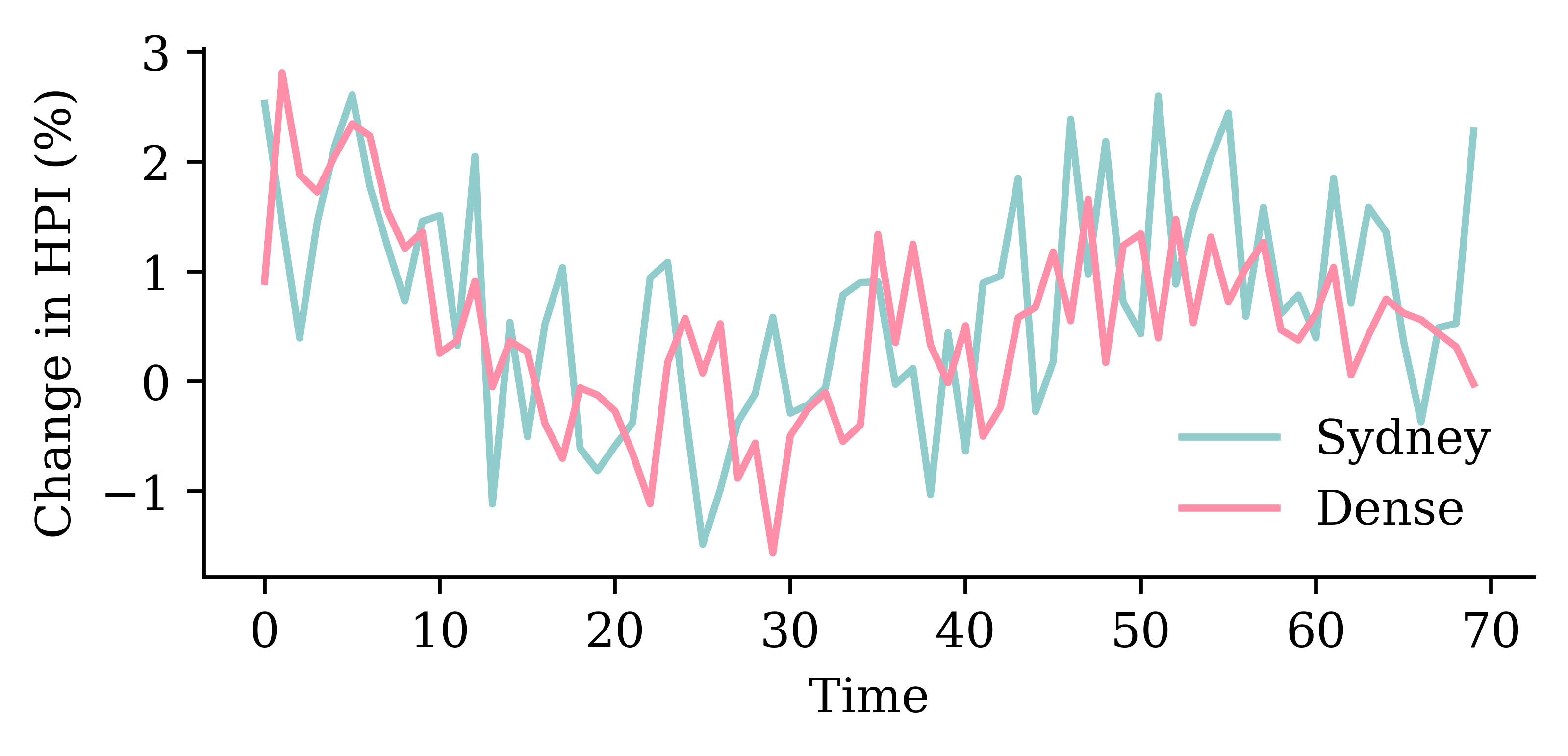
SimpleRNN layerrandom.seed(1)
model_simple = Sequential([
Input((seq_length, num_ts)),
SimpleRNN(50),
Dense(num_ts, activation="linear")
])
model_simple.compile(loss="mse", optimizer="adam")
print(f"This model has {model_simple.count_params()} parameters.")
es = EarlyStopping(patience=50, restore_best_weights=True, verbose=1)
%time hist = model_simple.fit(train_ds, epochs=1_000, \
validation_data=val_ds, callbacks=[es], verbose=0);This model has 3257 parameters.
Epoch 93: early stopping
Restoring model weights from the end of the best epoch: 43.
CPU times: user 5.08 s, sys: 2.59 s, total: 7.67 s
Wall time: 2.82 s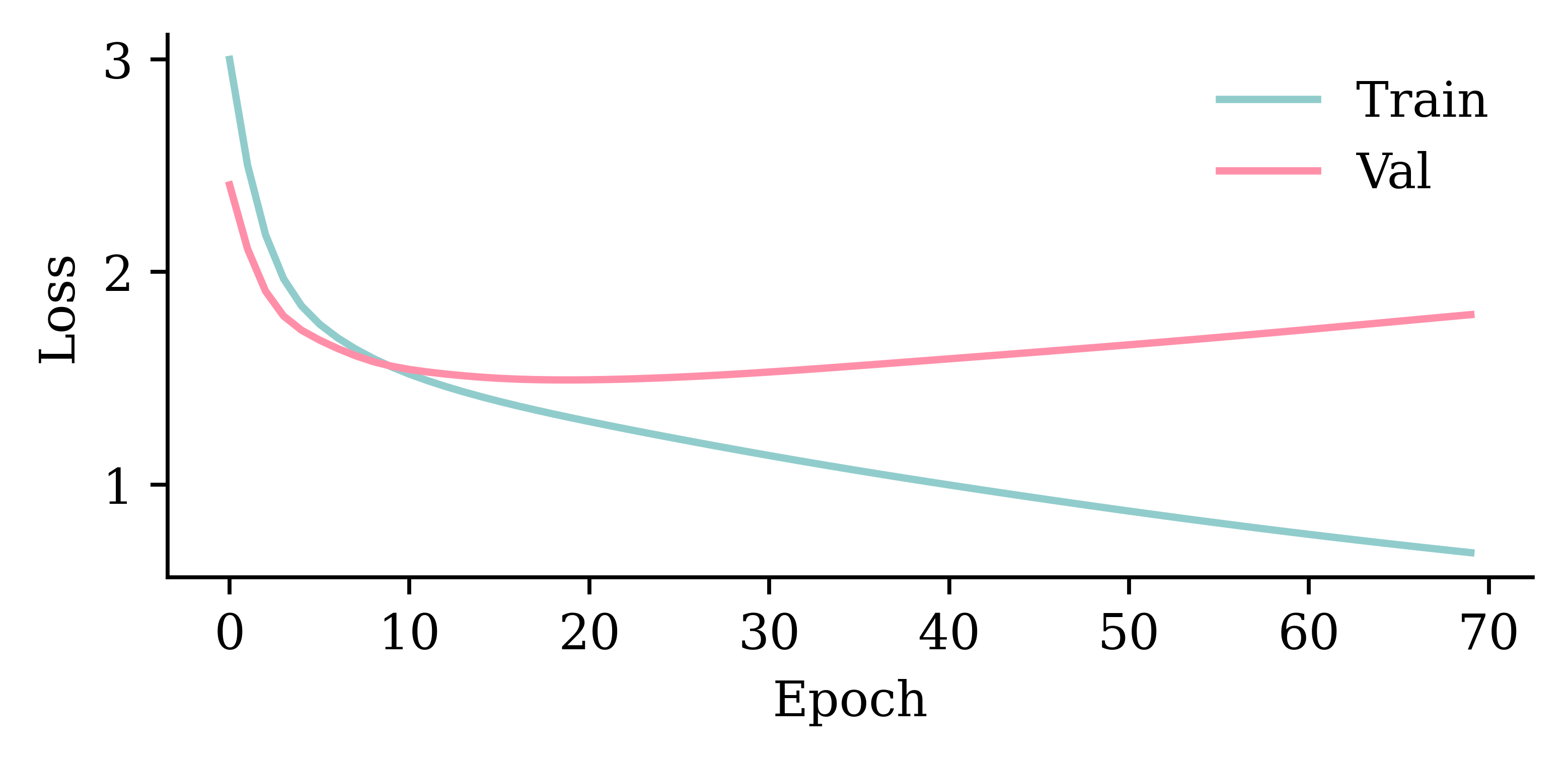
model_simple.evaluate(val_ds, verbose=0)1.6287589073181152plot_model(model_simple, show_shapes=True)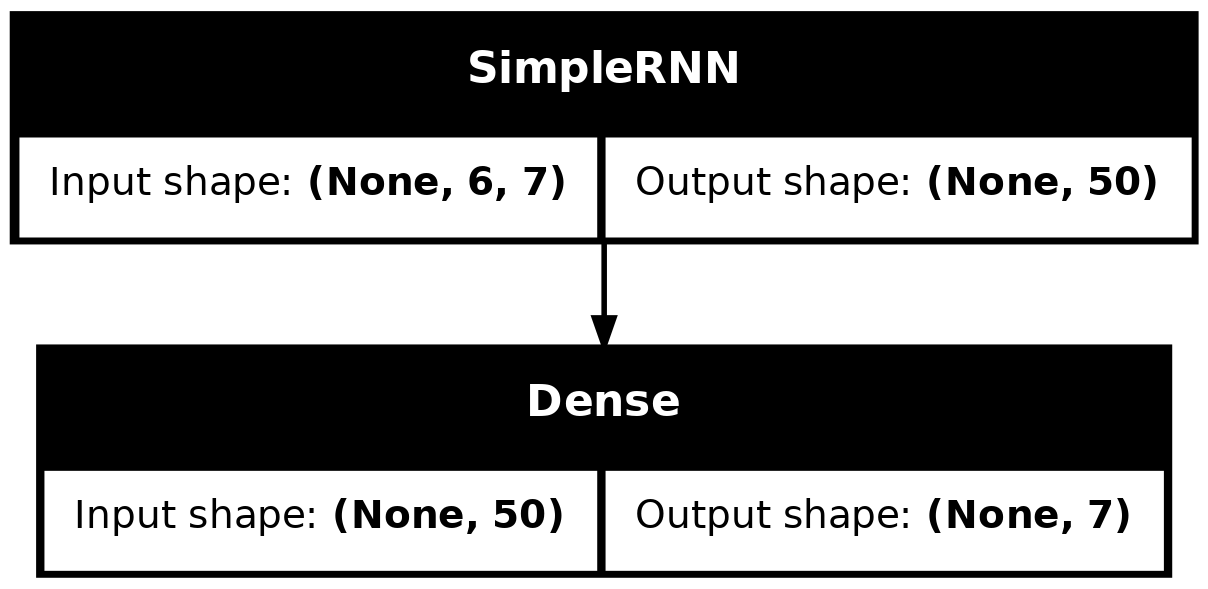
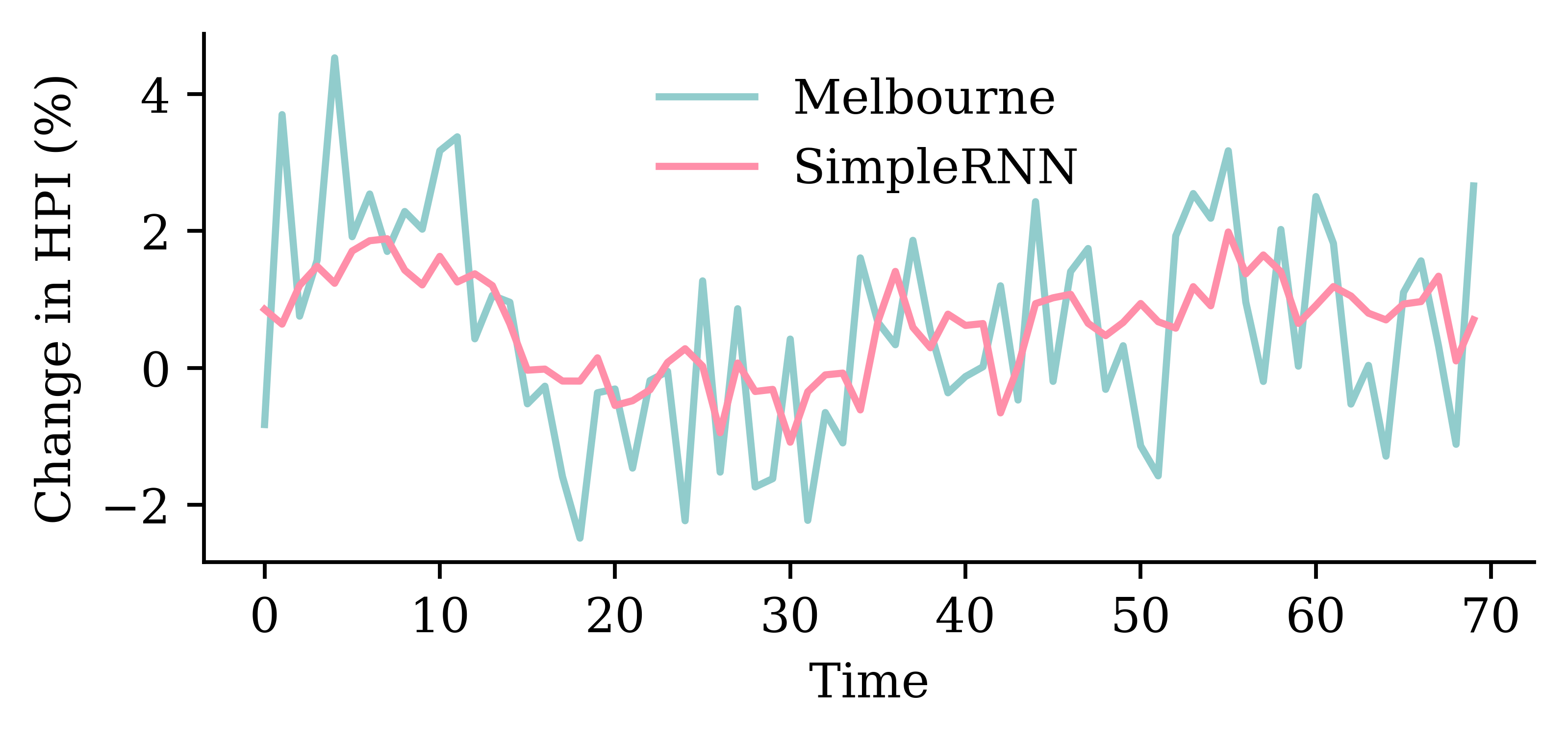

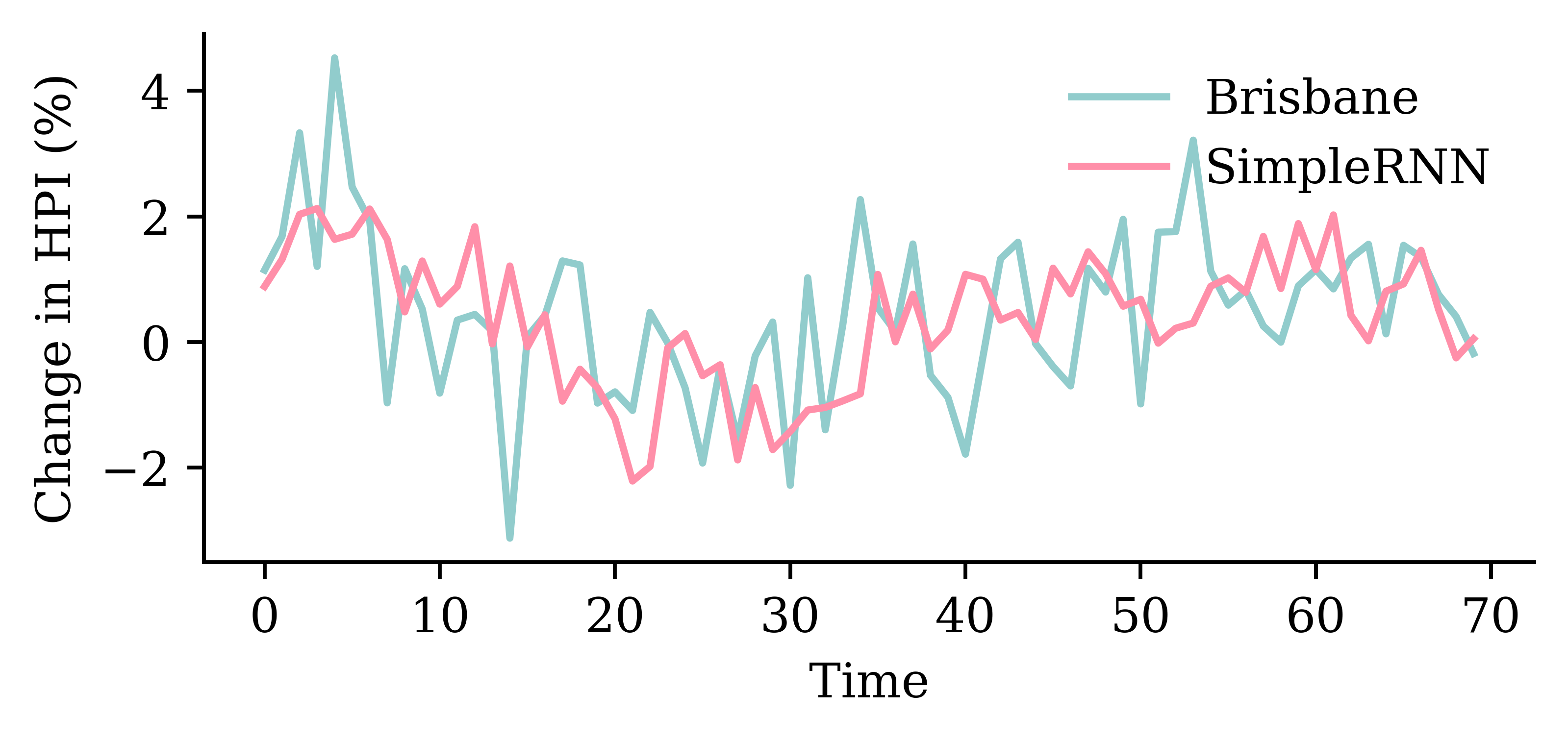
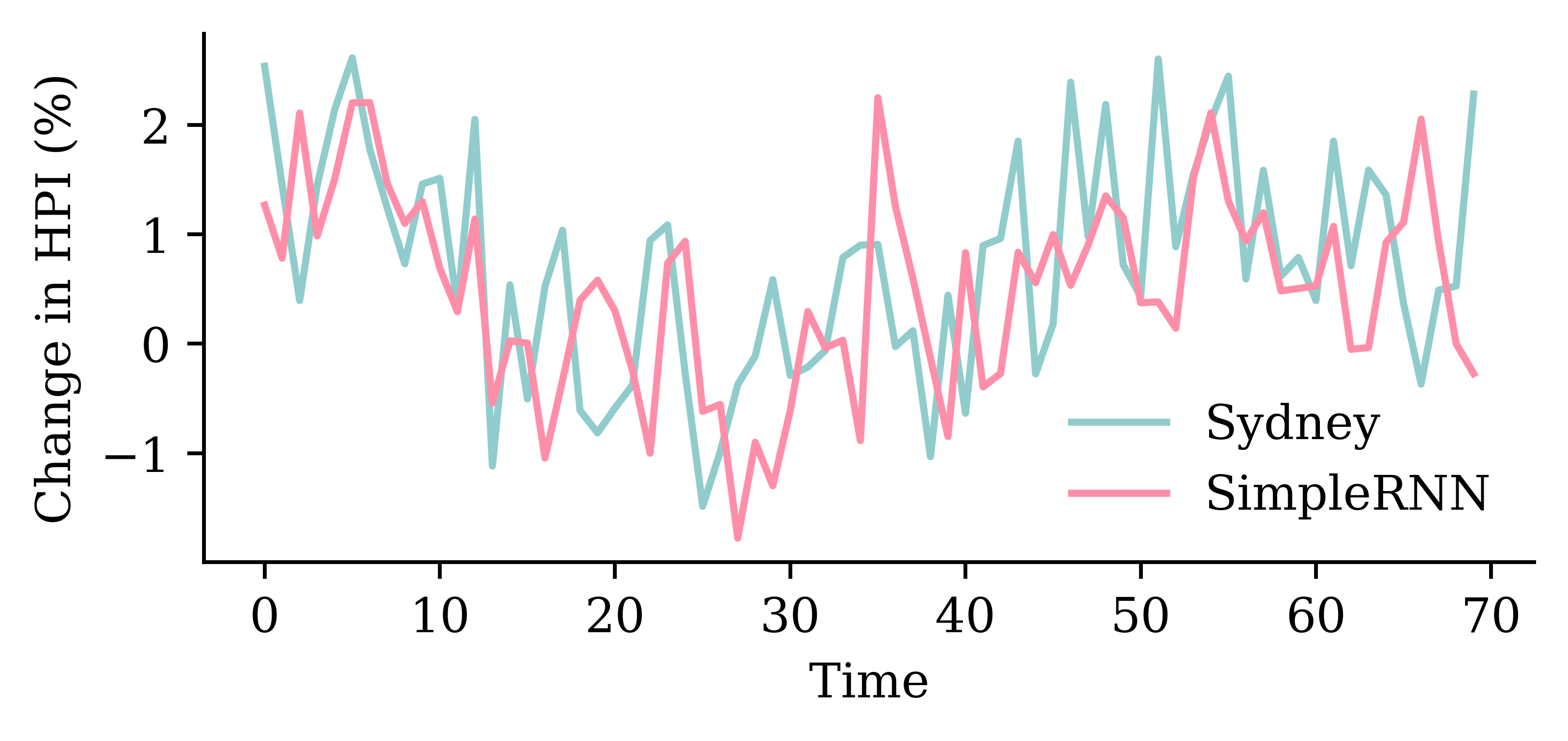
LSTM layerrandom.seed(1)
model_lstm = Sequential([
Input((seq_length, num_ts)),
LSTM(50),
Dense(num_ts, activation="linear")
])
model_lstm.compile(loss="mse", optimizer="adam")
es = EarlyStopping(patience=50, restore_best_weights=True, verbose=1)
%time hist = model_lstm.fit(train_ds, epochs=1_000, \
validation_data=val_ds, callbacks=[es], verbose=0);Epoch 106: early stopping
Restoring model weights from the end of the best epoch: 56.
CPU times: user 6.13 s, sys: 3.03 s, total: 9.16 s
Wall time: 3.56 s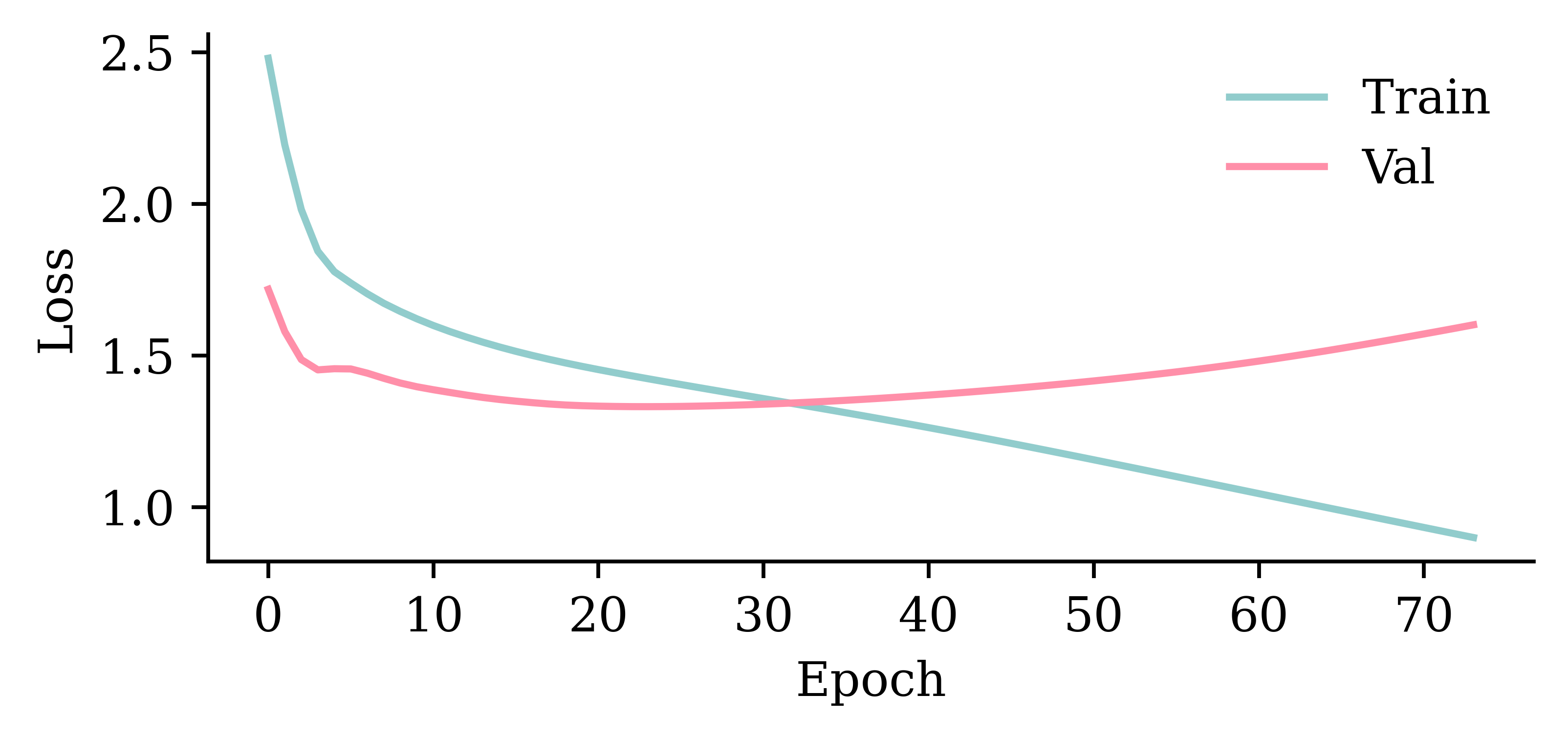
model_lstm.evaluate(val_ds, verbose=0)1.3439931869506836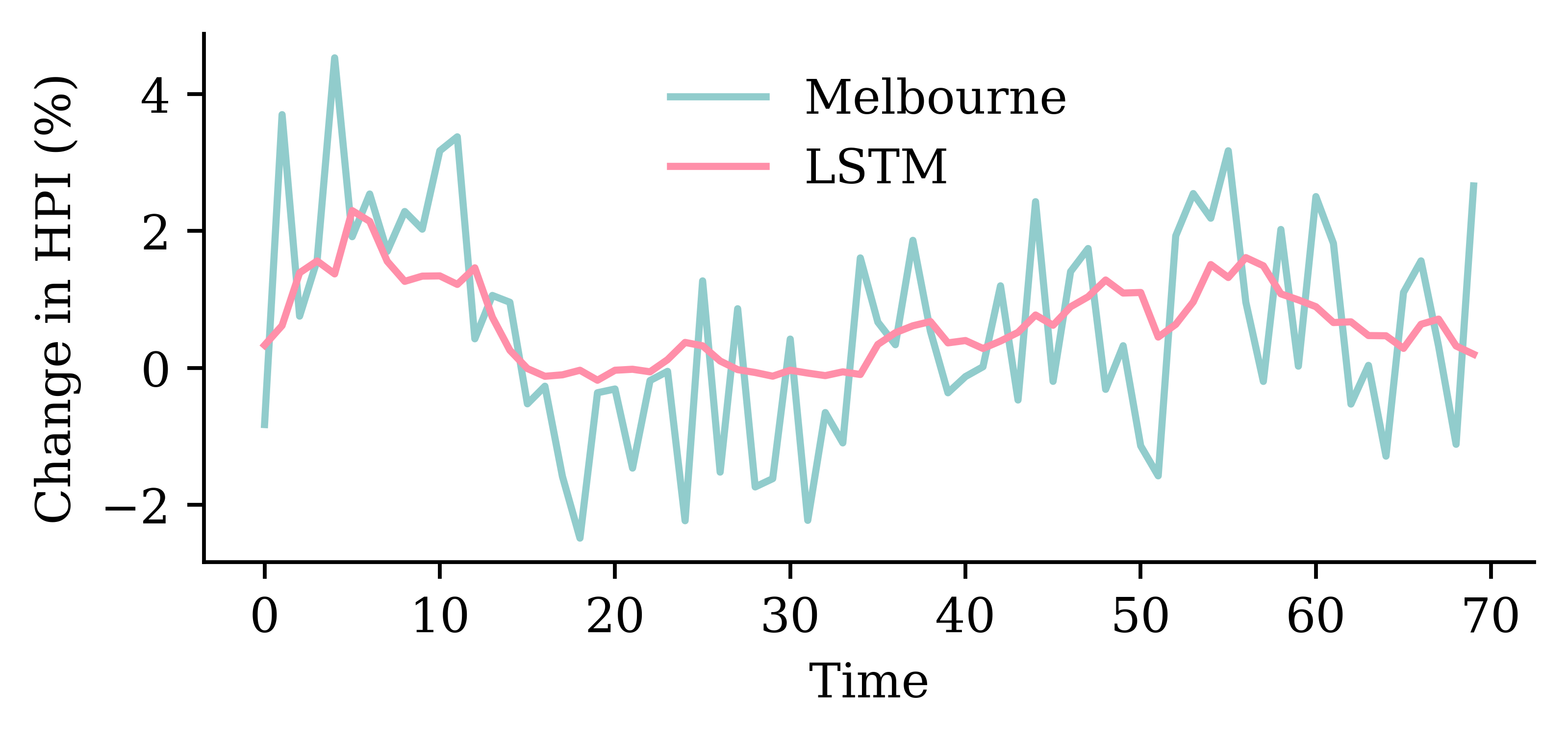

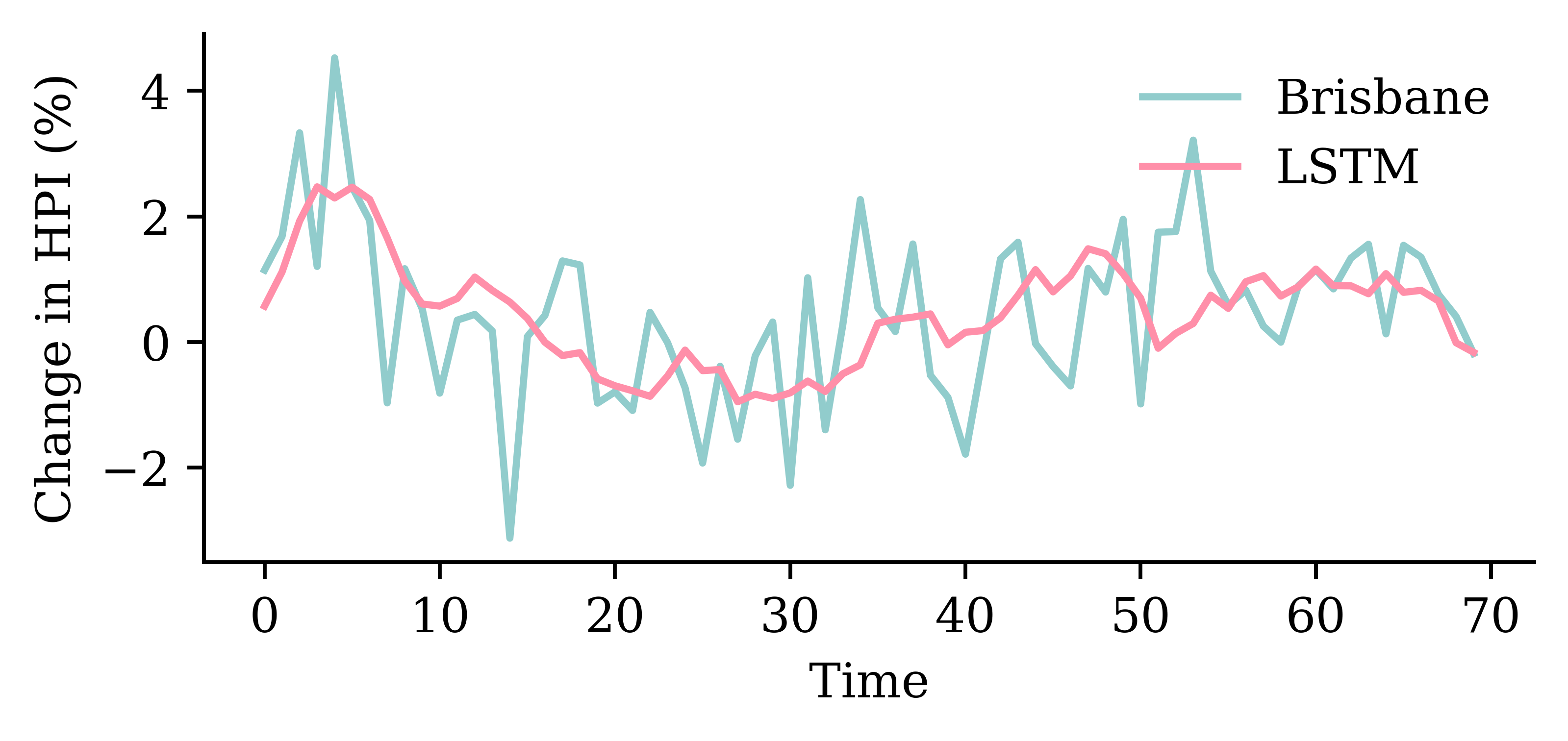
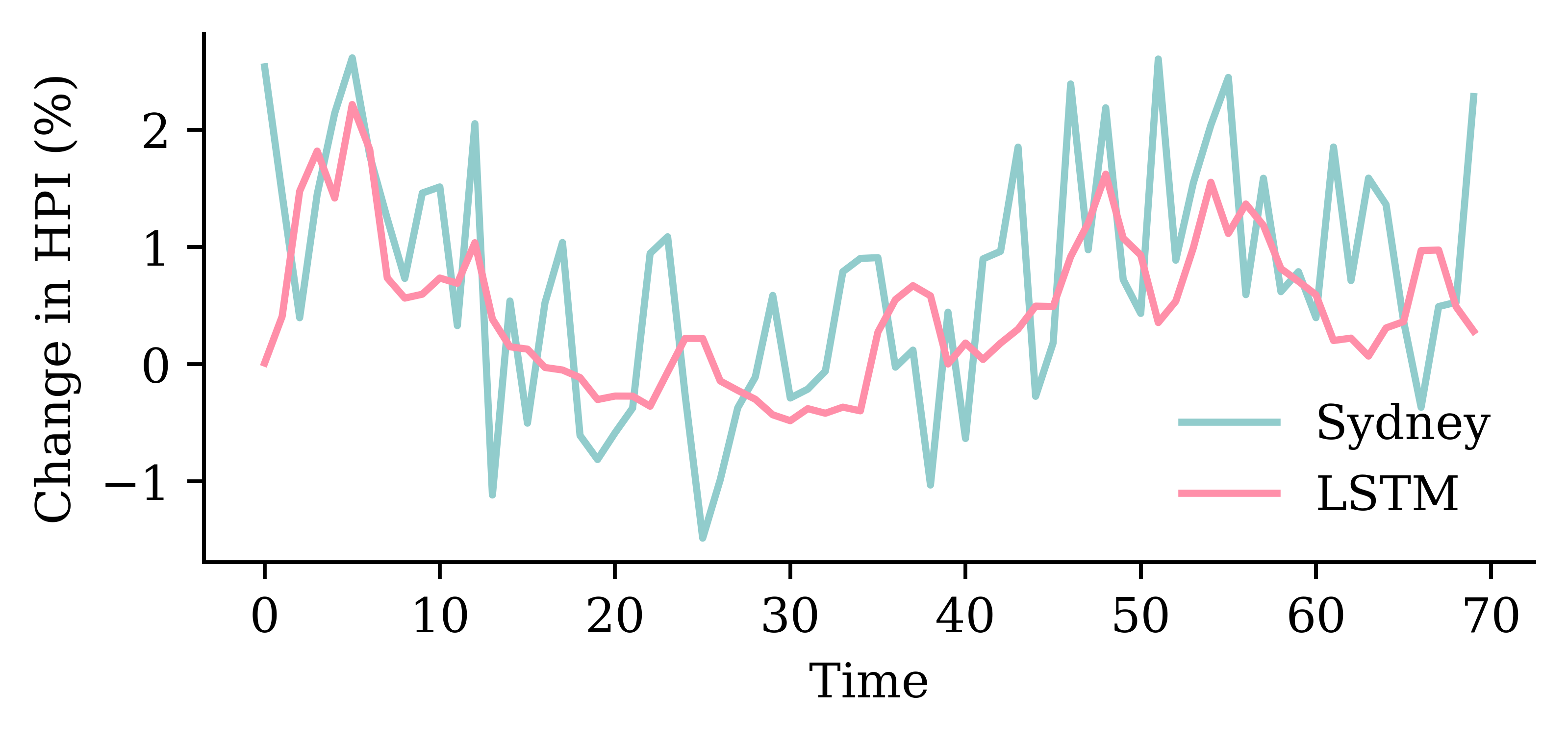
GRU layerrandom.seed(1)
model_gru = Sequential([
Input((seq_length, num_ts)),
GRU(50),
Dense(num_ts, activation="linear")
])
model_gru.compile(loss="mse", optimizer="adam")
es = EarlyStopping(patience=50, restore_best_weights=True, verbose=1)
%time hist = model_gru.fit(train_ds, epochs=1_000, \
validation_data=val_ds, callbacks=[es], verbose=0)Epoch 114: early stopping
Restoring model weights from the end of the best epoch: 64.
CPU times: user 6.62 s, sys: 3.21 s, total: 9.83 s
Wall time: 3.9 s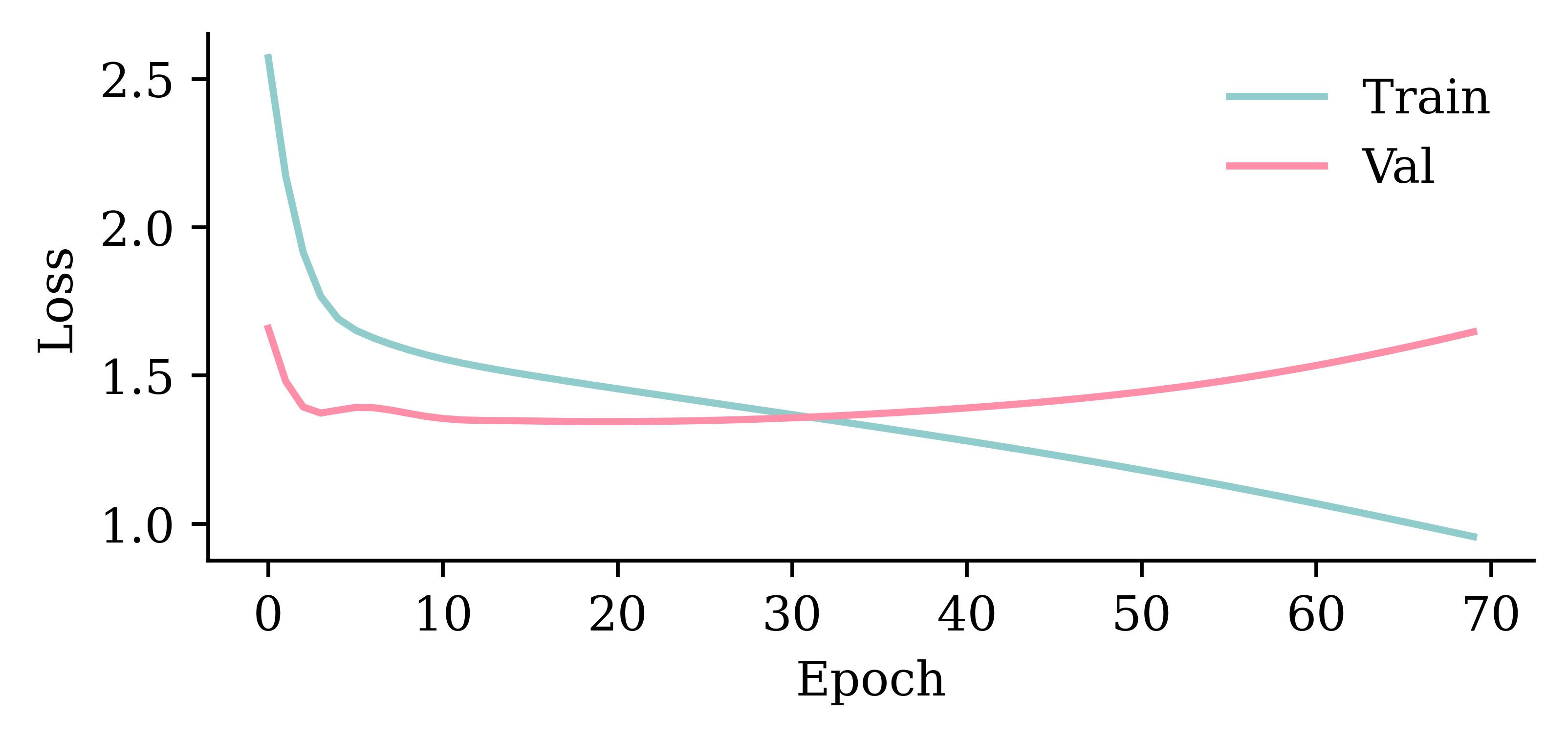
model_gru.evaluate(val_ds, verbose=0)1.3462404012680054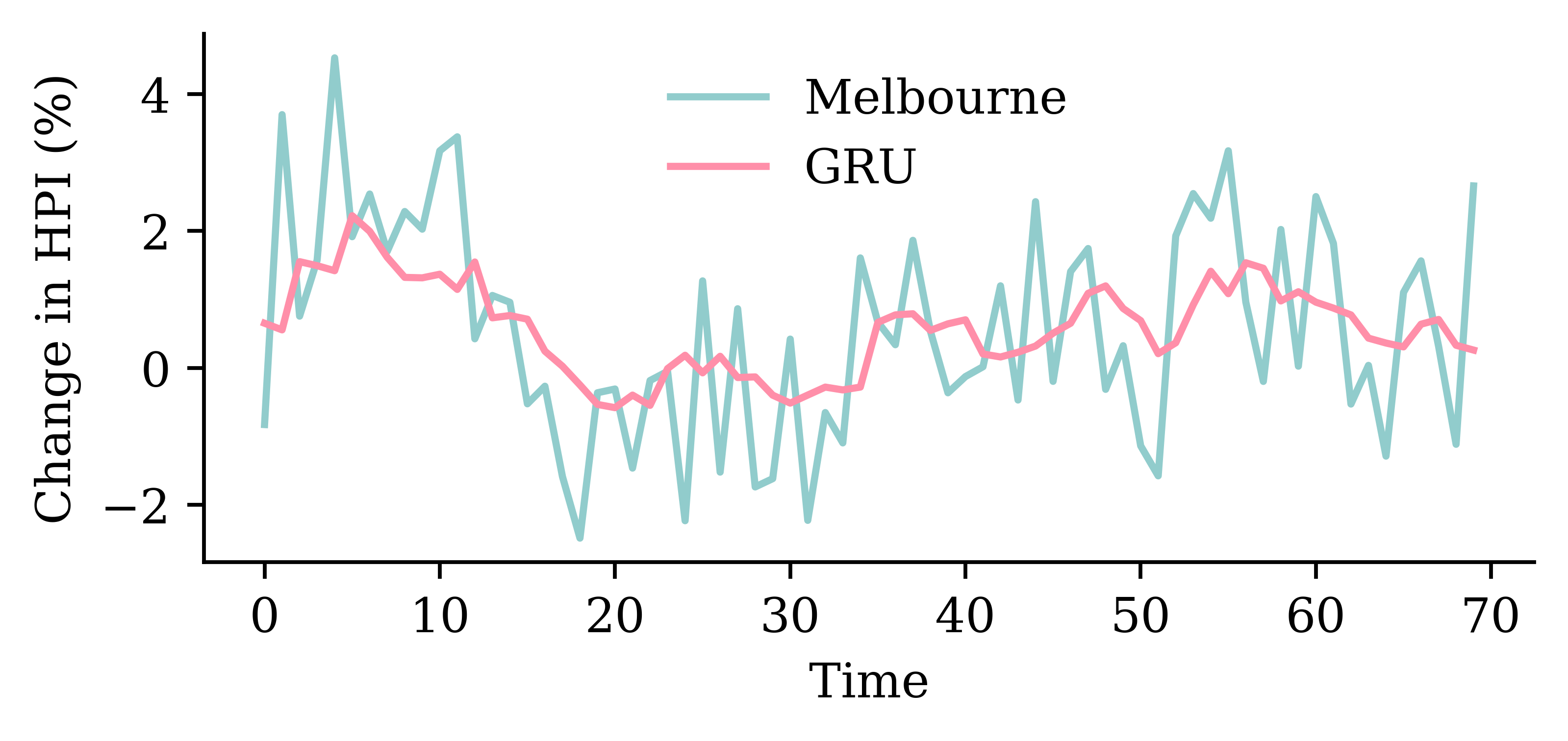

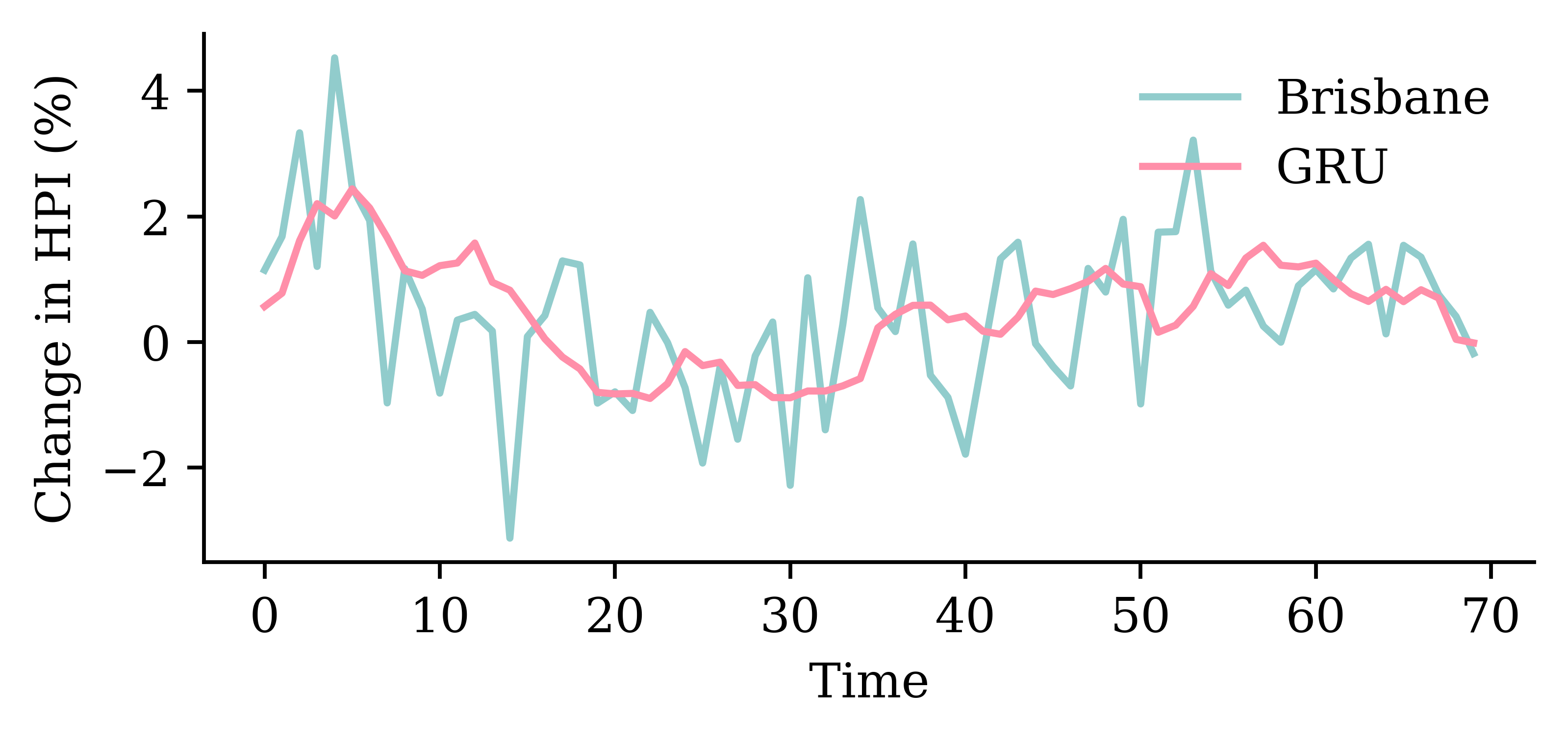
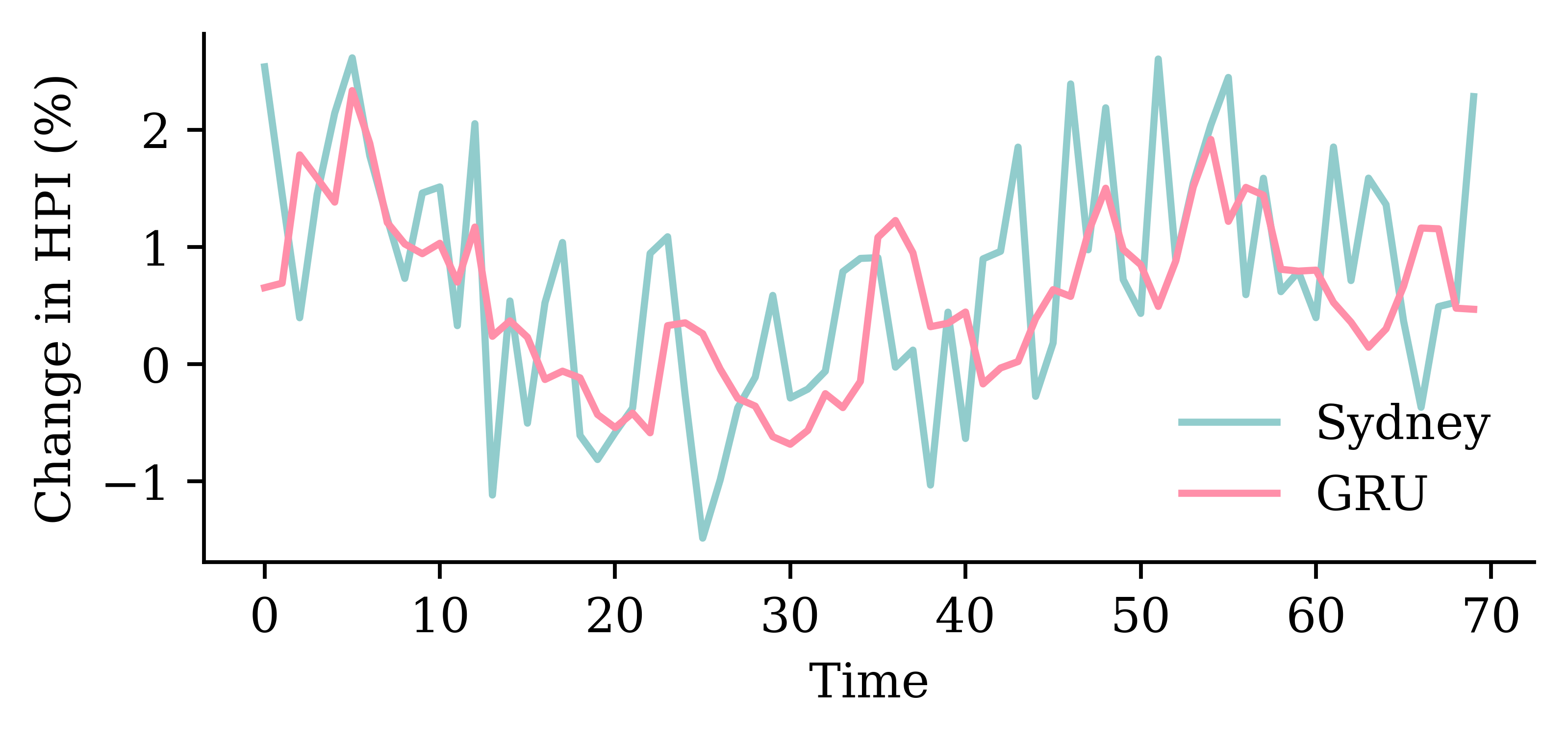
GRU layersrandom.seed(1)
model_two_grus = Sequential([
Input((seq_length, num_ts)),
GRU(50, return_sequences=True),
GRU(50),
Dense(num_ts, activation="linear")
])
model_two_grus.compile(loss="mse", optimizer="adam")
es = EarlyStopping(patience=50, restore_best_weights=True, verbose=1)
%time hist = model_two_grus.fit(train_ds, epochs=1_000, \
validation_data=val_ds, callbacks=[es], verbose=0)Epoch 93: early stopping
Restoring model weights from the end of the best epoch: 43.
CPU times: user 6.14 s, sys: 2.56 s, total: 8.71 s
Wall time: 3.93 s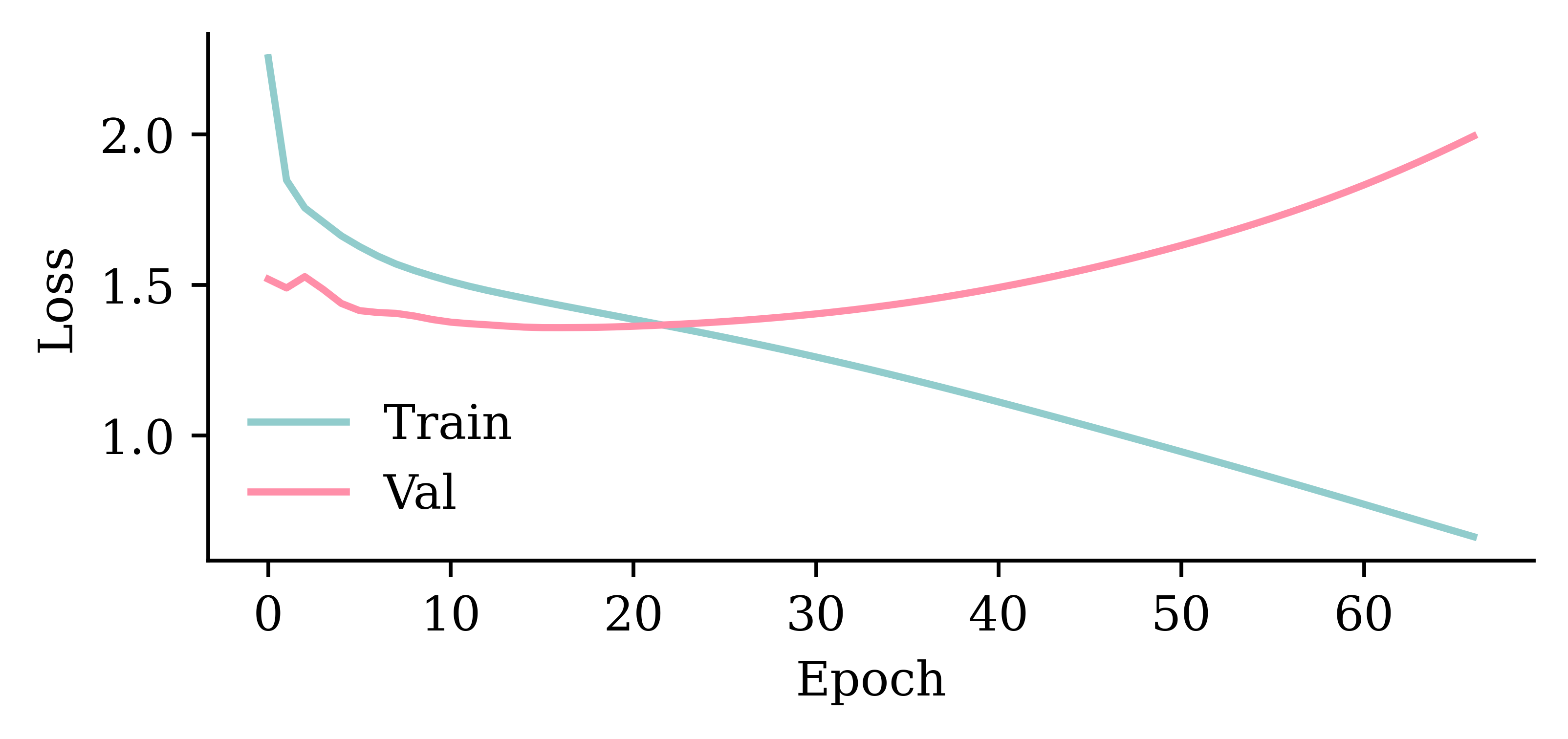
model_two_grus.evaluate(val_ds, verbose=0)1.3611823320388794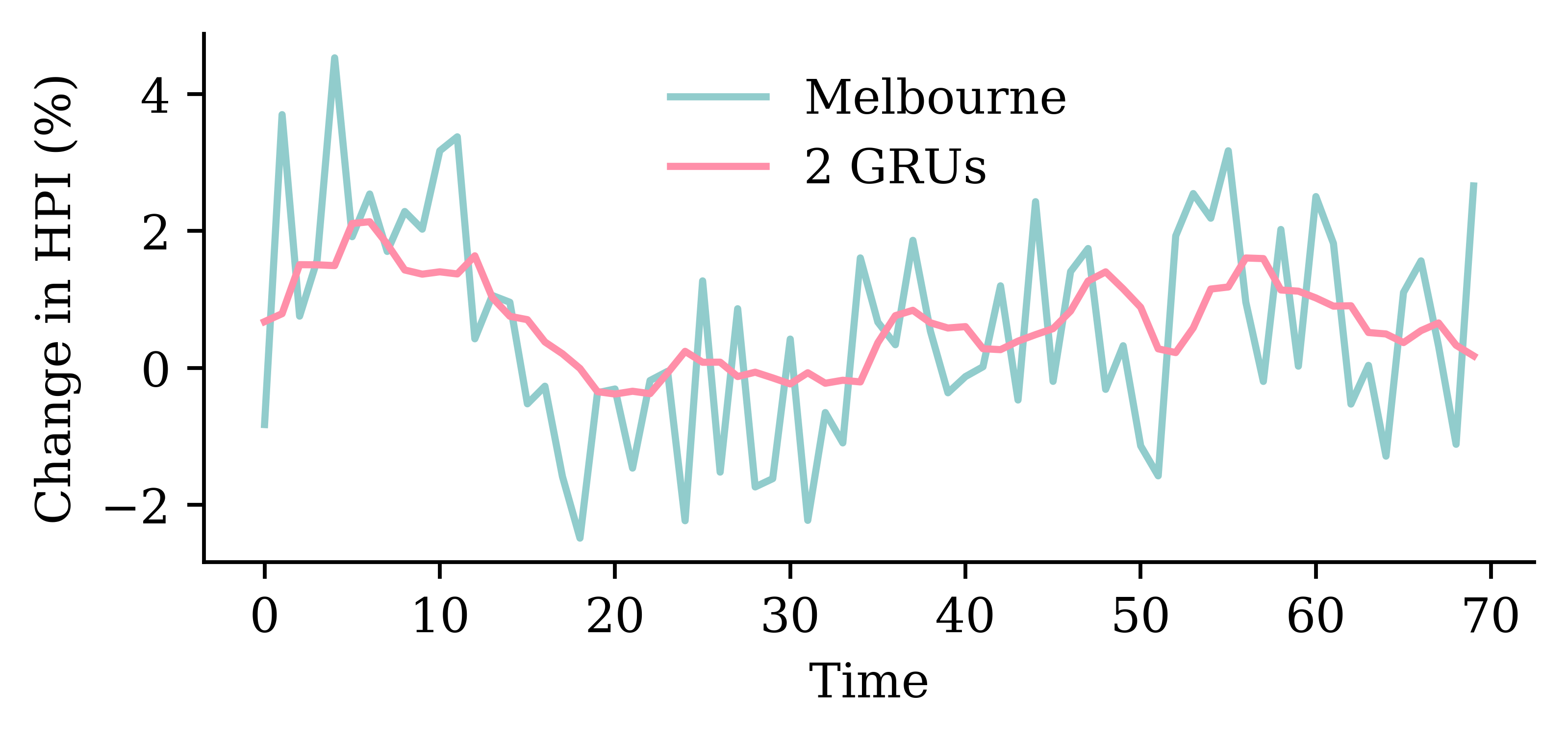

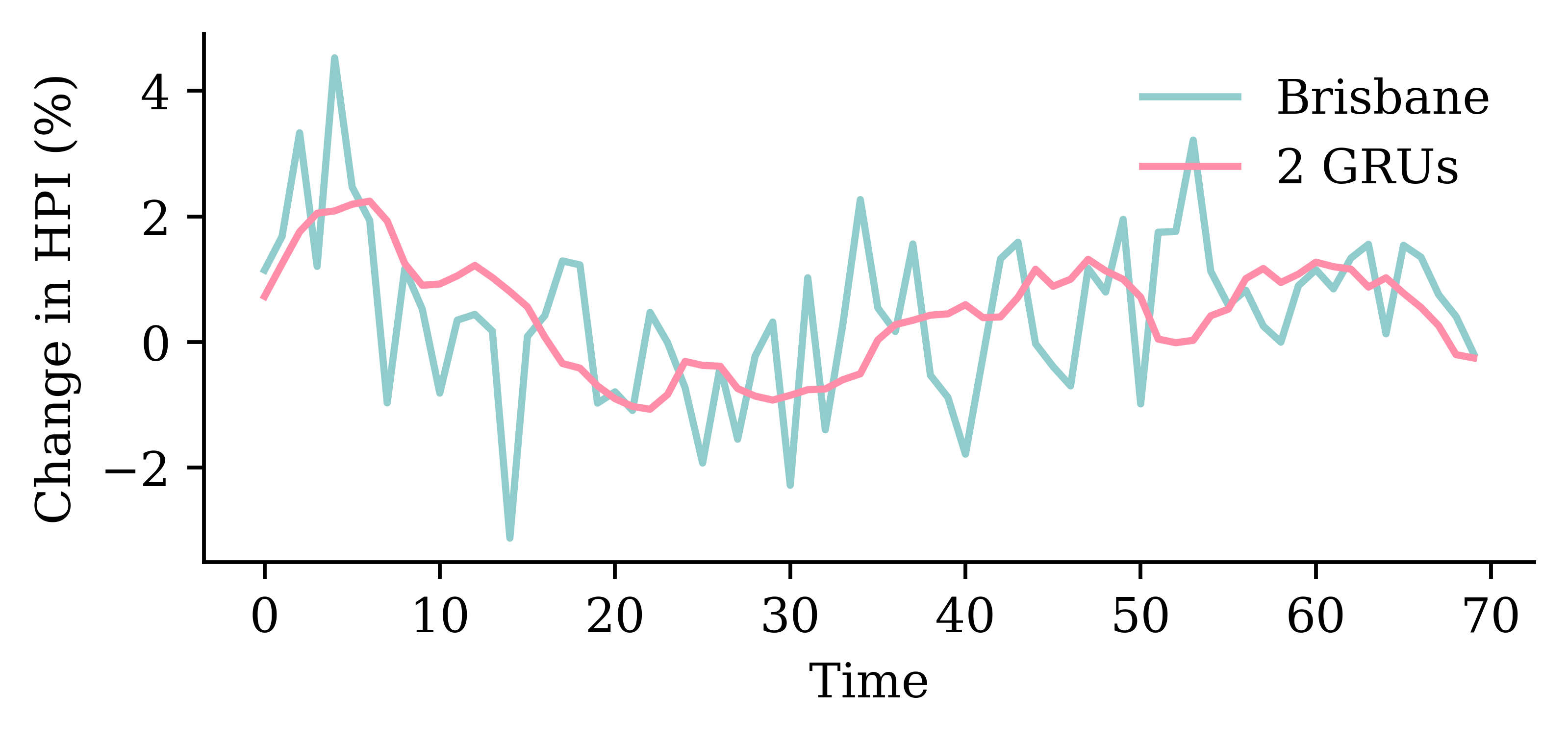
| Model | MSE | |
|---|---|---|
| 1 | SimpleRNN | 1.628759 |
| 0 | Dense | 1.445832 |
| 4 | 2 GRUs | 1.361182 |
| 3 | GRU | 1.346240 |
| 2 | LSTM | 1.343993 |
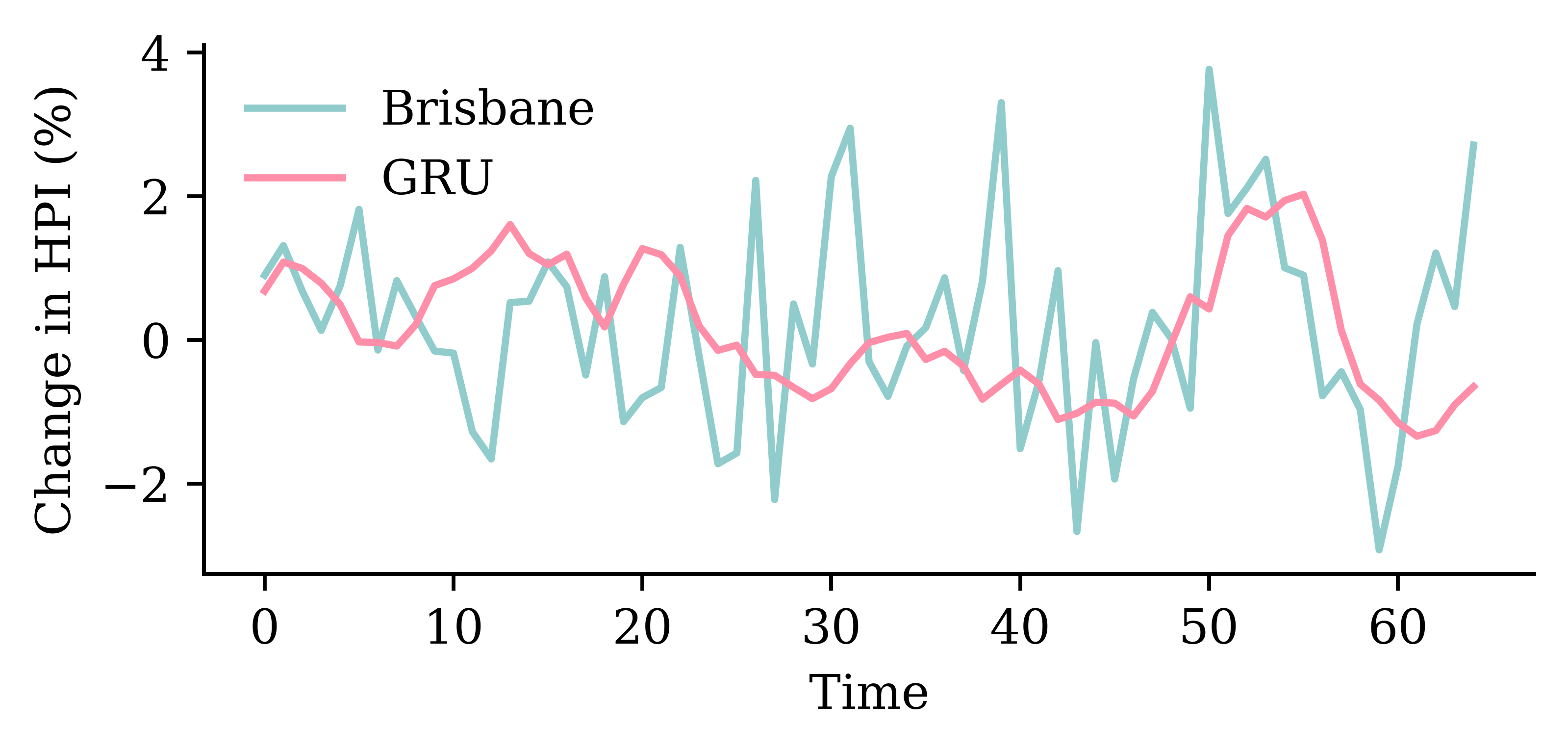
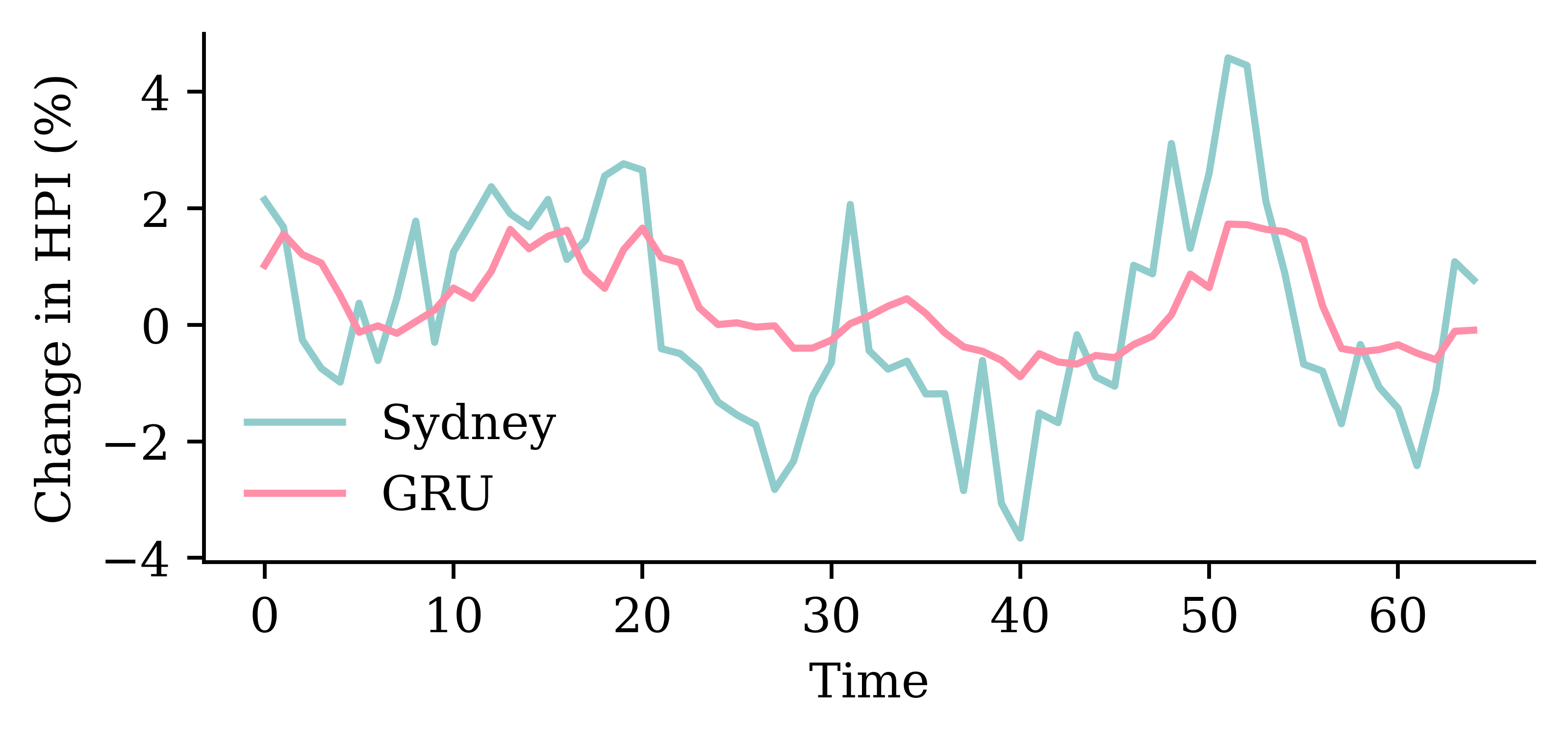
The network with an LSTM layer is the best.
model_lstm.evaluate(test_ds, verbose=0)1.879453420639038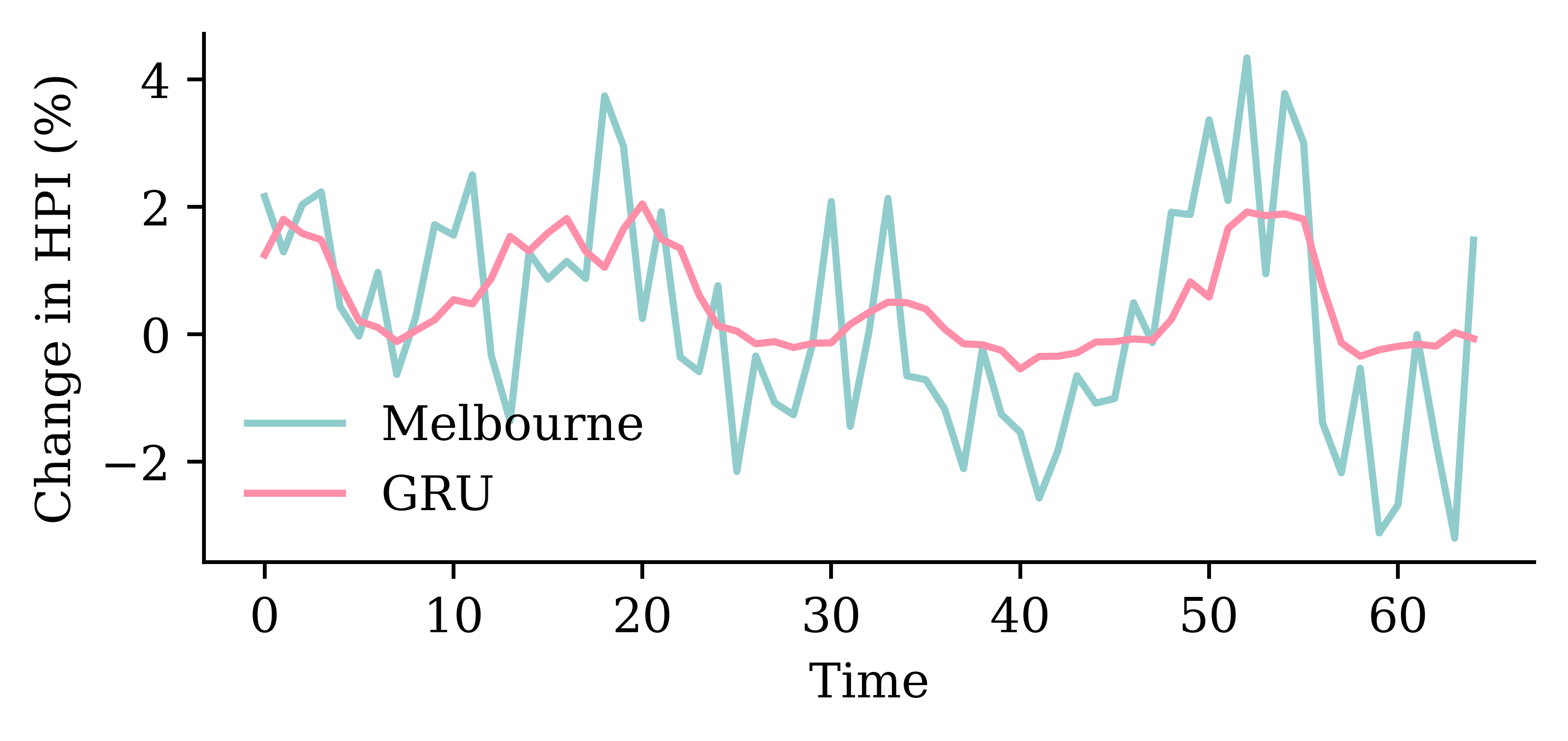

from watermark import watermark
print(watermark(python=True, packages="keras,matplotlib,numpy,pandas,seaborn,scipy,torch,tensorflow,tf_keras"))Python implementation: CPython
Python version : 3.11.8
IPython version : 8.23.0
keras : 3.2.0
matplotlib: 3.8.4
numpy : 1.26.4
pandas : 2.2.1
seaborn : 0.13.2
scipy : 1.11.0
torch : 2.2.2
tensorflow: 2.16.1
tf_keras : 2.16.0